C2Protein kinase C conserved region 2 (CalB) |
|---|
| SMART accession number: | SM00239 |
|---|---|
| Description: | Ca2+-binding motif present in phospholipases, protein kinases C, and synaptotagmins (among others). Some do not appear to contain Ca2+-binding sites. Particular C2s appear to bind phospholipids, inositol polyphosphates, and intracellular proteins. Unusual occurrence in perforin. Synaptotagmin and PLC C2s are permuted in sequence with respect to N- and C-terminal beta strands. SMART detects C2 domains using one or both of two profiles. |
| Interpro abstract (IPR000008): | The C2 domain is a Ca 2+ -dependent membrane-targeting module found in many cellular proteins involved in signal transduction or membrane trafficking. C2 domains are unique among membrane targeting domains in that they show wide range of lipid selectivity for the major components of cell membranes, including phosphatidylserine and phosphatidylcholine. This C2 domain is about 116 amino-acid residues and is located between the two copies of the C1 domain in Protein Kinase C and the protein kinase catalytic domain [ (PUBMED:22453964) ]. Regions with significant homology [ (PUBMED:7559667) ] to the C2-domain have been found in many proteins. The C2 domain is thought to be involved in calcium-dependent phospholipid binding [ (PUBMED:8253763) ] and in membrane targetting processes such as subcellular localisation. The 3D structure of the C2 domain of synaptotagmin has been reported [ (PUBMED:7697723) ], the domain forms an eight-stranded beta sandwich constructed around a conserved 4-stranded motif, designated a C2 key [ (PUBMED:7697723) ]. Calcium binds in a cup-shaped depression formed by the N- and C-terminal loops of the C2-key motif. Structural analyses of several C2 domains have shown them to consist of similar ternary structures in which three Ca 2+ -binding loops are located at the end of an 8 stranded antiparallel beta sandwich. |
| Family alignment: |
There are 169765 C2 domains in 106000 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing C2 domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with C2 domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing C2 domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Cellular role: signalling
Binding / catalysis: calcium-binding, phospholipid-binding, inositol polyphosphate-binding, membrane-binding - Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Rizo J, Sudhof TC
- C2-domains, structure and function of a universal Ca2+-binding domain.
- J Biol Chem. 1998; 273: 15879-82
- Hurley JH, Newton AC, Parker PJ, Blumberg PM, Nishizuka Y
- Taxonomy and function of C1 protein kinase C homology domains.
- Protein Sci. 1997; 6: 477-80
- Display abstract
C1 domains are compact alpha/beta structural units of about 50 amino acids which tightly bind two zinc ions. These domains were first discovered as the loci of phorbol ester and diacylglycerol binding to conventional protein kinase C isozymes, which contain 2 C1 domains (C1A and C1B) in their N-terminal regulatory regions. We present a comprehensive list of 54 C1 domains occurring singly or doubly in 34 different proteins. Many C1 domains and C1 domain-containing proteins bind phorbol esters, but many others do not. By combining analysis of 54 C1 domain sequences with information from previously reported solution and crystal structure determinations and site-directed mutagenesis, profiles are derived and used to classify C1 domains. Twenty-six C1 domains fit the profile for phorbol-ester binding and are termed "typical." Twenty-eight other domains fit the profile for the overall C1 domain fold but do not fit the profile for phorbol ester binding, and are termed "atypical." Proteins containing typical C1 domains are predicted to be regulated by diacylglycerol, whereas those containing only atypical domains are not.
- Puls A, Schmidt S, Grawe F, Stabel S
- Interaction of protein kinase C zeta with ZIP, a novel protein kinase C-binding protein.
- Proc Natl Acad Sci U S A. 1997; 94: 6191-6
- Display abstract
The atypical protein kinase C (PKC) member PKC-zeta has been implicated in several signal transduction pathways regulating differentiation, proliferation or apoptosis of mammalian cells. We report here the identification of a cytoplasmic and membrane-associated protein that we name zeta-interacting protein (ZIP) and that interacts with the regulatory domain of PKC-zeta but not classic PKCs. The structural motifs in ZIP include a recently defined ZZ zinc finger as a potential protein binding module, two PEST sequences and a novel putative protein binding motif with the consensus sequence YXDEDX5SDEE/D. ZIP binds to the pseudosubstrate region in the regulatory domain of PKC-zeta and is phosphorylated by PKC-zeta in vitro. ZIP dimerizes via the same region that promotes binding to PKC-zeta suggesting a competitive situation between ZIP:ZIP and ZIP:PKC-zeta complexes. In the absence of PKC-zeta proper subcellular localization of ZIP is impaired and we show that intracellular targeting of ZIP is dependent on a balanced interaction with PKC-zeta. Taking into account the recent isolation of ZIP by others in different contexts we propose that ZIP may function as a scaffold protein linking PKC-zeta to protein tyrosine kinases and cytokine receptors.
- Nalefski EA, Falke JJ
- The C2 domain calcium-binding motif: structural and functional diversity.
- Protein Sci. 1996; 5: 2375-90
- Display abstract
The C2 domain is a Ca(2+)-binding motif of approximately 130 residues in length originally identified in the Ca(2+)-dependent isoforms of protein kinase C. Single and multiple copies of C2 domains have been identified in a growing number of eukaryotic signalling proteins that interact with cellular membranes and mediate a broad array of critical intracellular processes, including membrane trafficking, the generation of lipid-second messengers, activation of GTPases, and the control of protein phosphorylation. As a group, C2 domains display the remarkable property of binding a variety of different ligands and substrates, including Ca2+, phospholipids, inositol polyphosphates, and intracellular proteins. Expanding this functional diversity is the fact that not all proteins containing C2 domains are regulated by Ca2+, suggesting that some C2 domains may play a purely structural role or may have lost the ability to bind Ca2+. The present review summarizes the information currently available regarding the structure and function of the C2 domain and provides a novel sequence alignment of 65 C2 domain primary structures. This alignment predicts that C2 domains form two distinct topological folds, illustrated by the recent crystal structures of C2 domains from synaptotagmin 1 and phosphoinositide-specific phospholipase C-delta 1, respectively. The alignment highlights residues that may be critical to the C2 domain fold or required for Ca2+ binding and regulation.
- Ponting CP, Parker PJ
- Extending the C2 domain family: C2s in PKCs delta, epsilon, eta, theta, phospholipases, GAPs, and perforin.
- Protein Sci. 1996; 5: 162-6
- Display abstract
Various membrane lipid metabolites, generated by phospholipases C and D (PLCs, PLDs), are known to regulate the activities of protein kinases C (PKCs) and GTP-ase activating proteins (GAPs) in a range of cellular processes. Conventional Ca(2+)-dependent PKCs (alpha, beta I, beta II, and gamma), PLCs and various GAPs are all known to contain copies of a phospholipid-binding domain, termed C2 or CalB. Here we recognize that C2 domains are also present in "new" Ca(2+)-independent PKCs (delta, epsilon, eta, and theta), other kinases, a eukaryotic PLD, the breakpoint cluster region (BCR) gene product, and two further GAPS. Twenty-two previously unrecognized C2 domain sequences are presented, which include a single copy in the mammalian poreforming proteins, perforin.
- Shao X, Davletov BA, Sutton RB, Sudhof TC, Rizo J
- Bipartite Ca2+-binding motif in C2 domains of synaptotagmin and protein kinase C.
- Science. 1996; 273: 248-51
- Display abstract
C2 domains are found in many proteins involved in membrane traffic or signal transduction. Although C2 domains are thought to bind calcium ions, the structural basis for calcium binding is unclear. Analysis of calcium binding to C2 domains of synaptotagmin I and protein kinase C-beta by nuclear magnetic resonance spectroscopy revealed a bipartite calcium-binding motif that involves the coordination of two calcium ions by five aspartate residues located on two separate loops. Sequence comparisons indicated that this may be a widely used calcium-binding motif, designated here as the C2 motif.
- Li C, Ullrich B, Zhang JZ, Anderson RG, Brose N, Sudhof TC
- Ca(2+)-dependent and -independent activities of neural and non-neural synaptotagmins.
- Nature. 1995; 375: 594-9
- Display abstract
Synaptotagmins (Syts) are brain-specific Ca2+/phospholipid-binding proteins. In hippocampal synapses, Syt I is essential for fast Ca(2+)-dependent synaptic vesicle exocytosis but not for Ca(2+)-independent exocytosis. In vertebrates and invertebrates, Syt may therefore participate in Ca(2+)-dependent synaptic membrane fusion, either by serving as the Ca2+ sensor in the last step of fast Ca(2+)-triggered neurotransmitter release, or by collaborating with an additional Ca2+ sensor. While Syt I binds Ca2+ (refs 10, 11), its phospholipid binding is triggered at lower calcium concentrations (EC50 = 3-6 microM) than those required for exocytosis. Furthermore, Syts bind clathrin-AP2 with high affinity, indicating that they may play a general role in endocytosis rather than being confined to a specialized function in regulated exocytosis. Here we resolve this apparent contradiction by describing four Syts, three of which (Syt VI, VII and VIII) are widely expressed in non-neural tissues. All Syts tested share a common domain structure, with a cytoplasmic region composed of two C2 domains that interacts with clathrin-AP2 (Kd = 0.1-1.0 nM) and with neural and non-neural syntaxins. The first C2 domains of Syt I, II, III, V and VII, but not of IV, VI or VIII, bind phospholipids with a similar Ca(2+)-concentration dependence (EC50 = 3-6 microM). The same C2 domains also bind syntaxin as a function of Ca2+ but the Ca(2+)-concentration dependence of Syt I, II and V (> 200 microM) differs from that of Syt III and VII (< 10 microM).(ABSTRACT TRUNCATED AT 250 WORDS)
- Sutton RB, Davletov BA, Berghuis AM, Sudhof TC, Sprang SR
- Structure of the first C2 domain of synaptotagmin I: a novel Ca2+/phospholipid-binding fold.
- Cell. 1995; 80: 929-38
- Display abstract
C2 domains are regulatory sequence motifs that occur widely in nature. Synaptotagmin I, a synaptic vesicle protein involved in the Ca2+ regulation of exocytosis, contains two C2 domains, the first of which acts as a Ca2+ sensor. We now describe the three-dimensional structure of this C2 domain at 1.9 A resolution in both the Ca(2+)-bound and Ca(2+)-free forms. The C2 polypeptide forms an eight-stranded beta sandwich constructed around a conserved four-stranded motif designated as a C2 key. Ca2+ binds in a cup-shaped depression between two polypeptide loops located at the N- and C-termini of the C2-key motif.
- Brose N, Petrenko AG, Sudhof TC, Jahn R
- Synaptotagmin: a calcium sensor on the synaptic vesicle surface.
- Science. 1992; 256: 1021-5
- Display abstract
Neurons release neurotransmitters by calcium-dependent exocytosis of synaptic vesicles. However, the molecular steps transducing the calcium signal into membrane fusion are still an enigma. It is reported here that synaptotagmin, a highly conserved synaptic vesicle protein, binds calcium at physiological concentrations in a complex with negatively charged phospholipids. This binding is specific for calcium and involves the cytoplasmic domain of synaptotagmin. Calcium binding is dependent on the intact oligomeric structure of synaptotagmin (it is abolished by proteolytic cleavage at a single site). These results suggest that synaptotagmin acts as a cooperative calcium receptor in exocytosis.
- Perin MS, Fried VA, Mignery GA, Jahn R, Sudhof TC
- Phospholipid binding by a synaptic vesicle protein homologous to the regulatory region of protein kinase C.
- Nature. 1990; 345: 260-3
- Display abstract
Neurotransmitters are released at synapses by the Ca2(+)-regulated exocytosis of synaptic vesicles, which are specialized secretory organelles that store high concentrations of neurotransmitters. The rapid Ca2(+)-triggered fusion of synaptic vesicles is presumably mediated by specific proteins that must interact with Ca2+ and the phospholipid bilayer. We now report that the cytoplasmic domain of p65, a synaptic vesicle-specific protein that binds calmodulin contains an internally repeated sequence that is homologous to the regulatory C2-region of protein kinase C (PKC). The cytoplasmic domain of recombinant p65 binds acidic phospholipids with a specificity indicating an interaction of p65 with the hydrophobic core as well as the headgroups of the phospholipids. The binding specificity resembles PKC, except that p65 also binds calmodulin, placing the C2-regions in a context of potential Ca2(+)-regulation that is different from PKC. This is a novel homology between a cellular protein and the regulatory domain of protein kinase C. The structure and properties of p65 suggest that it may have a role in mediating membrane interactions during synaptic vesicle exocytosis.
- Disease (disease genes where sequence variants are found in this domain)
-
SwissProt sequences and OMIM curated human diseases associated with missense mutations within the C2 domain.
Protein Disease Perforin-1 (P14222) (SMART) OMIM:170280: Hemophagocytic lymphohistiocytosis, familial, 2
OMIM:603553: - Metabolism (metabolic pathways involving proteins which contain this domain)
-
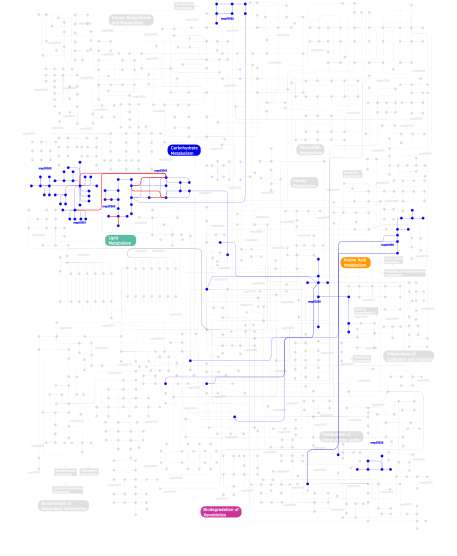
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 10.12 map04070 Phosphatidylinositol signaling system 7.26  map00562
map00562Inositol phosphate metabolism 7.19 map04020 Calcium signaling pathway 4.46 map04730 Long-term depression 3.93 map04370 VEGF signaling pathway 3.93 map04650 Natural killer cell mediated cytotoxicity 3.86 map04310 Wnt signaling pathway 3.86 map04720 Long-term potentiation 3.86 map04916 Melanogenesis 3.86 map04540 Gap junction 3.60 map04010 MAPK signaling pathway 3.33 map04670 Leukocyte transendothelial migration 3.33 map05214 Glioma 3.33 map05223 Non-small cell lung cancer 3.33 map04012 ErbB signaling pathway 3.13 map04530 Tight junction 2.93 map04912 GnRH signaling pathway 2.33 map04510 Focal adhesion 2.00 map04664 Fc epsilon RI signaling pathway 1.60 map04120 Ubiquitin mediated proteolysis 1.53  map00564
map00564Glycerophospholipid metabolism 1.13 map04620 Toll-like receptor signaling pathway 1.00 map05120 Epithelial cell signaling in Helicobacter pylori infection 1.00  map00565
map00565Ether lipid metabolism 0.87 map05050 Dentatorubropallidoluysian atrophy (DRPLA) 0.80 map05220 Chronic myeloid leukemia 0.73 map04350 TGF-beta signaling pathway 0.73 map04360 Axon guidance 0.60 map04140 Regulation of autophagy 0.60  map00590
map00590Arachidonic acid metabolism 0.60 map00592 alpha-Linolenic acid metabolism 0.60 map04940 Type I diabetes mellitus 0.60  map00591
map00591Linoleic acid metabolism 0.53 map05040 Huntington's disease 0.47  map00260
map00260Glycine, serine and threonine metabolism 0.40 map05222 Small cell lung cancer 0.40 map05213 Endometrial cancer 0.40 map04662 B cell receptor signaling pathway 0.40 map05212 Pancreatic cancer 0.40 map04630 Jak-STAT signaling pathway 0.40 map04810 Regulation of actin cytoskeleton 0.40 map05210 Colorectal cancer 0.40 map04210 Apoptosis 0.40 map04910 Insulin signaling pathway 0.40 map05215 Prostate cancer 0.40 map04150 mTOR signaling pathway 0.40 map04660 T cell receptor signaling pathway 0.40 map05211 Renal cell carcinoma 0.40 map05221 Acute myeloid leukemia 0.40 map04930 Type II diabetes mellitus 0.40 map05218 Melanoma 0.40 map04914 Progesterone-mediated oocyte maturation 0.13  map00380
map00380Tryptophan metabolism 0.07  map00632
map00632Benzoate degradation via CoA ligation This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with C2 domain which could be assigned to a KEGG orthologous group, and not all proteins containing C2 domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of C2 domains in PDB
PDB code Main view Title 1a25 
C2 DOMAIN FROM PROTEIN KINASE C (BETA) 1bci 
C2 DOMAIN OF CYTOSOLIC PHOSPHOLIPASE A2, NMR, MINIMIZED AVERAGE STRUCTURE 1bdy 
C2 DOMAIN FROM PROTEIN KINASE C DELTA 1byn 
SOLUTION STRUCTURE OF THE CALCIUM-BOUND FIRST C2-DOMAIN OF SYNAPTOTAGMIN I 1cjy 
HUMAN CYTOSOLIC PHOSPHOLIPASE A2 1djg 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH LANTHANUM 1djh 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH BARIUM 1dji 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH CALCIUM 1djw 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH INOSITOL-2-METHYLENE-1,2-CYCLIC-MONOPHOSPHONATE 1djx 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH INOSITOL-1,4,5-TRISPHOSPHATE 1djy 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH INOSITOL-2,4,5-TRISPHOSPHATE 1djz 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH INOSITOL-4,5-BISPHOSPHATE 1dqv 
CRYSTAL STRUCTURE OF SYNAPTOTAGMIN III C2A/C2B 1dsy 
C2 DOMAIN FROM PROTEIN KINASE C (ALPHA) COMPLEXED WITH CA2+ AND PHOSPHATIDYLSERINE 1gmi 
Structure of the c2 domain from novel protein kinase C epsilon 1k5w 
THREE-DIMENSIONAL STRUCTURE OF THE SYNAPTOTAGMIN 1 C2B-DOMAIN: SYNAPTOTAGMIN 1 AS A PHOSPHOLIPID BINDING MACHINE 1qas 
1-PHOSPHATIDYLINOSITOL-4,5-BISPHOSPHATE PHOSPHODIESTERASE DELTA 1 1qat 
1-PHOSPHATIDYLINOSITOL-4,5-BISPHOSPHATE PHOSPHODIESTERASE DELTA COMPLEX WITH SAMARIUM (III) CHLORIDE 1rh8 
Three-dimensional structure of the calcium-free Piccolo C2A-domain 1rlw 
CALCIUM-PHOSPHOLIPID BINDING DOMAIN FROM CYTOSOLIC PHOSPHOLIPASE A2 1rsy 
STRUCTURE OF THE FIRST C2-DOMAIN OF SYNAPTOTAGMIN I: A NOVEL CA2+(SLASH)PHOSPHOLIPID BINDING FOLD 1tjm 
Crystallographic Identification of Sr2+ Coordination Site in Synaptotagmin I C2B Domain 1tjx 
Crystallographic Identification of Ca2+ Coordination Sites in Synaptotagmin I C2B Domain 1ugk 
Solution structure of the first C2 domain of synaptotagmin IV from human fetal brain (KIAA1342) 1uov 
Calcium binding domain C2B 1uow 
Calcium binding domain C2B 1v27 
Solution structure of the first C2 domain of RIM2 1w15 
rat synaptotagmin 4 C2B domain in the presence of calcium 1w16 
rat synaptotagmin 4 C2B domain in the absence of calcium 1wfj 
C2 domain-containing protein from putative elicitor-responsive gene 1yrk 
The C2 Domain of PKC is a new Phospho-Tyrosine Binding Domain 2b3r 
Crystal structure of the C2 domain of class II phosphatidylinositide 3-kinase C2 2bwq 
Crystal Structure of the RIM2 C2A-domain at 1.4 angstrom Resolution 2chd 
Crystal structure of the C2A domain of Rabphilin-3A 2cjs 
Structural Basis for a Munc13-1 Homodimer - Munc13-1 - RIM Heterodimer Switch: C2-domains as Versatile Protein-Protein Interaction Modules 2cjt 
Structural Basis for a Munc13-1 Homodimer - Munc13-1 - RIM Heterodimer Switch: C2-domains as Versatile Protein-Protein Interaction Modules 2cm5 
crystal structure of the C2B domain of rabphilin 2cm6 
crystal structure of the C2B domain of rabphilin3A 2d8k 
Solution structure of the first C2 domain of synaptotagmin VII 2dmg 
Solution structure of the third C2 domain of KIAA1228 protein 2dmh 
Solution structure of the first C2 domain of human myoferlin 2enp 
Solution structure of the first C2 domain from human B/K protein 2enq 
Solution structure of the C2 domain from human PI3-kinase p110 subunit alpha 2ep6 
Solution structure of the second C2 domain from human MCTP2 protein 2fju 
Activated Rac1 bound to its effector phospholipase C beta 2 2fk9 
Human protein kinase C, eta 2isd 
PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT 2jqz 
Solution Structure of the C2 domain of human Smurf2 2k3h 
Structural determinants for Ca2+ and PIP2 binding by the C2A domain of rabphilin-3A 2k45 
C2A domain of synaptototagmin I solution structure in the FGF-1-C2A binary complex: key component in the fibroblast growthfactor non-classical pathway 2k4a 
FGF-1-C2A binary complex structure: a key component in the fibroblast growthfactor non-classical pathway 2k8m 
S100A13-C2A binary complex structure 2ki6 
The FGF1-S100A13-C2A hetero-hexameric complex structure: A component in the non-classical pathway for FGF1 secretion 2lha 
Solution structure of C2B with IP6 2n1t 
2N1T 2nq3 
Crystal structure of the C2 Domain of Human Itchy Homolog E3 Ubiquitin Protein Ligase 2nsq 
Crystal structure of the C2 domain of the human E3 ubiquitin-protein ligase NEDD4-like protein 2q3x 
The RIM1alpha C2B domain 2r83 
Crystal structure analysis of human synaptotagmin 1 C2A-C2B 2rd0 
Structure of a human p110alpha/p85alpha complex 2uzp 
Crystal structure of the C2 domain of human protein kinase C gamma. 2yoa 
Synaptotagmin-1 C2B domain with phosphoserine 2zkm 
Crystal Structure of Phospholipase C Beta 2 3b7y 
Crystal structure of the C2 Domain of the E3 Ubiquitin-Protein Ligase NEDD4 3bxj 
Crystal Structure of the C2-GAP Fragment of synGAP 3f00 
Crystal Structure of Synaptotagmin I C2A domain with Cu(II) 3f01 
Crystal Structure of Synaptotagmin I C2A domain with Cu(II) 3f04 
Crystal Structure of Synaptotagmin I C2A domain 3f05 
Crystal Structure of Synaptotagmin I C2A domain with Mn(II) 3fbk 
Crystal structure of the C2 domain of the human regulator of G-protein signaling 3 isoform 6 (RGP3), Northeast Structural Genomics Consortium Target HR5550A 3fdw 
Crystal structure of a C2 domain from human synaptotagmin-like protein 4 3gpe 
Crystal Structure Analysis of PKC (alpha)-C2 domain complexed with Ca2+ and PtdIns(4,5)P2 3hhm 
Crystal structure of p110alpha H1047R mutant in complex with niSH2 of p85alpha and the drug wortmannin 3hiz 
Crystal structure of p110alpha H1047R mutant in complex with niSH2 of p85alpha 3hn8 
Crystal structure of synaptotagmin 3jzy 
Crystal structure of human Intersectin 2 C2 domain 3kwt 
Munc13-1 C2B-domain, calcium-free 3kwu 
Munc13-1 C2B-domain, calcium bound 3l9b 
Crystal Structure of Rat Otoferlin C2A 3m7f 
Crystal structure of the Nedd4 C2/Grb10 SH2 complex 3n5a 
Synaptotagmin-7, C2B-domain, calcium bound 3nsj 
The X-ray crystal structure of lymphocyte perforin 3ohm 
Crystal structure of activated G alpha Q bound to its effector phospholipase C beta 3 3pfq 
Crystal Structure and Allosteric Activation of Protein Kinase C beta II 3pyc 
Crystal structure of human SMURF1 C2 domain 3qr0 
Crystal Structure of S. officinalis PLC21 3qr1 
Crystal Structure of L. pealei PLC21 3rdj 
Rat PKC C2 domain Apo 3rpb 
THE C2B-DOMAIN OF RABPHILIN: STRUCTURAL VARIATIONS IN A JANUS-FACED DOMAIN 3twy 
RAT PKC C2 DOMAIN BOUND TO PB 3w56 
Structure of a C2 domain 3w57 
Structure of a C2 domain 3zim 
Discovery of a potent and isoform-selective targeted covalent inhibitor of the lipid kinase PI3Kalpha 4a55 
Crystal structure of p110alpha in complex with iSH2 of p85alpha and the inhibitor PIK-108 4dnl 
Crystal structure of a C2 domain of a protein kinase C alpha (PRKCA) from Homo sapiens at 1.90 A resolution 4gnk 
Crystal structure of Galphaq in complex with full-length human PLCbeta3 4icw 
4ICW 4icx 
4ICX 4ihb 
X-RAY STRUCTURE OF THE canonical C2A DOMAIN FROM HUMAN DYSFERLIN 4iqh 
Crystal Structure Analysis of Dysferlin C2A variant 1 (C2Av1) 4jps 
Co-crystal Structures of the Lipid Kinase PI3K alpha with Pan and Isoform Selective Inhibitors 4l1b 
Crystal Structure of p110alpha complexed with niSH2 of p85alpha 4l1l 
Rat PKC C2 domain bound to CD 4l23 
Crystal Structure of p110alpha complexed with niSH2 of p85alpha and PI-103 4l2y 
Crystal Structure of p110alpha complexed with niSH2 of p85alpha and compound 9d 4lcv 
Crystal Structure of DOC2B C2A domain 4ldc 
Crystal Structure of DOC2B C2B domain 4lt7 
Crystal structure of the c2a domain of rabphilin-3a in complex with a calcium 4mjj 
Crystal structure of the C2A domain of DOC2A 4np9 
Structure of Rabphilin C2A domain bound to IP3 4npj 
Extended-Synaptotagmin 2, C2A- and C2B-domains 4npk 
Extended-Synaptotagmin 2, C2A- and C2B-domains, calcium bound 4ns0 
The C2A domain of Rabphilin 3A in complex with PI(4,5)P2 4ovu 
4OVU 4ovv 
4OVV 4p42 
Extended-Synaptotagmin 2, SMP - C2A - C2B Domains 4qj3 
4QJ3 4qj4 
4QJ4 4qj5 
4QJ5 4rj9 
4RJ9 4ts6 
4TS6 4tuu 
4TUU 4tv3 
4TV3 4v11 
4V11 4v29 
4V29 4waf 
4WAF 4wee 
4WEE 4y1s 
4Y1S 4y1t 
4Y1T 4ykn 
4YKN 4zop 
4ZOP 5a4x 
5A4X 5a50 
5A50 5a51 
5A51 5a52 
5A52 5ccg 
5CCG 5cch 
5CCH 5cci 
5CCI 5ccj 
5CCJ 5dxh 
5DXH 5dxt 
5DXT 5fi4 
5FI4 5h4y 
5H4Y 5h4z 
5H4Z 5itd 
5ITD 5ixc 
5IXC 5iz5 
5IZ5 5izr 
5IZR 5kj7 
5KJ7 5kj8 
5KJ8 - Links (links to other resources describing this domain)
-
PROSITE C2_DOMAIN_2 INTERPRO IPR000008 PFAM C2

