The domain within your query sequence starts at position and ends at position ; the E-value for the AT_hook domain shown below is < 1e-12.
dummy
AT_hookDNA binding domain with preference for A/T rich regions |
|---|
| SMART accession number: | SM00384 |
|---|---|
| Description: | Small DNA-binding motif first described in the high mobility group non-histone chromosomal protein HMG-I(Y). |
| Interpro abstract (IPR017956): | AT hooks are DNA-binding motifs with a preference for A/T rich regions. These motifs are found in a variety of proteins, including the high mobility group (HMG) proteins [ (PUBMED:11406267) ], in DNA-binding proteins from plants [ (PUBMED:8790293) ] and in hBRG1 protein, a central ATPase of the human switching/sucrose non-fermenting (SWI/SNF) remodeling complex [ (PUBMED:17081121) ]. |
| GO function: | DNA binding (GO:0003677) |
| Family alignment: |
There are 76954 AT_hook domains in 42187 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing AT_hook domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with AT_hook domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing AT_hook domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Binding / catalysis: DNA-binding
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Chuang RY, Kelly TJ
- The fission yeast homologue of Orc4p binds to replication origin DNA via multiple AT-hooks.
- Proc Natl Acad Sci U S A. 1999; 96: 2656-61
- Display abstract
The origin recognition complex (ORC) was originally identified in the yeast Saccharomyces cerevisiae as a protein that specifically binds to origins of DNA replication. Although ORC appears to play an essential role in the initiation of DNA replication in the cells of all eukaryotes, its interactions with DNA have not been defined in species other than budding yeast. We have characterized a Schizosaccharomyces pombe homologue of the ORC subunit, Orc4p. The homologue (Orp4p) consists of two distinct functional domains. The C-terminal domain shows strong sequence similarity to human, frog, and yeast Orc4 proteins, including conserved ATP-binding motifs. The N-terminal domain contains nine copies of the AT-hook motif found in a number of DNA-binding proteins, including the members of the HMG-I(Y) family of chromatin proteins. AT-hook motifs are known from biochemical and structural studies to mediate binding to the minor groove of AT-tracts in DNA. Orp4p is essential for viability of Sc. pombe and is expressed throughout the cell cycle. The Orp4 protein (and its isolated N-terminal domain) binds to the Sc. pombe replication origin, ars1. The DNA binding properties of Orp4p provide a plausible explanation for the characteristic features of Sc. pombe origins of replication, which differ significantly from those of Sa. cerevisiae.
- Aravind L, Landsman D
- AT-hook motifs identified in a wide variety of DNA-binding proteins.
- Nucleic Acids Res. 1998; 26: 4413-21
- Display abstract
The AT-hook is a small DNA-binding protein motif which was first described in the high mobility group non-histone chromosomal protein HMG-I(Y). Since its discovery, this motif has been observed in other DNA-binding proteins from a wide range of organisms. Using pattern searches and position-dependent matrices, we have extracted the AT-hook motifs present in a non-redundant protein sequence database. We have classified these motifs into three types according to their sequence similarity and have found that they are prevalent in many eukaryotic nuclear proteins in single or multiple copies. Furthermore, AT-hook motifs are frequently associated with known functional domains seen in chromatin proteins and in DNA-binding proteins (e.g. histone folds, homeodomains and zinc fingers). In general, it appears that the AT-hook motif is an auxiliary protein motif cooperating with other DNA-binding activities and facilitating changes in the structure of the DNA either as a polypeptide on its own [e.g. HMG-I(Y)] or as part of a multidomain protein [e.g. Swi2p in Saccharomyces cerevisiae or HRX (ALL-1) in Homo sapiens]. It is most interesting that this motif seems to be quite specific to known or predicted chromosomal/DNA-binding proteins, suggesting that it may act as a versatile minor groove tether.
- Huth JR et al.
- The solution structure of an HMG-I(Y)-DNA complex defines a new architectural minor groove binding motif.
- Nat Struct Biol. 1997; 4: 657-65
- Display abstract
The solution structure of a complex between a truncated form of HMG-I(Y), consisting of the second and third DNA binding domains (residues 51-90), and a DNA dodecamer containing the PRDII site of the interferon-beta promoter has been solved by multidimensional nuclear magnetic resonance spectroscopy. The stoichiometry of the complex is one molecule of HMG-I(Y) to two molecules of DNA. The structure reveals a new architectural minor groove binding motif which stabilizes B-DNA, thereby facilitating the binding of other transcription factors in the opposing major groove. The interactions involve a central Arg-Gly-Arg motif together with two other modules that participate in extensive hydrophobic and polar contracts. The absence of one of these modules in the third DNA binding domain accounts for its-100 fold reduced affinity relative to the second one.
- Todd RB, Andrianopoulos A
- Evolution of a fungal regulatory gene family: the Zn(II)2Cys6 binuclear cluster DNA binding motif.
- Fungal Genet Biol. 1997; 21: 388-405
- Display abstract
The coevolution of DNA binding proteins and their cognate binding sites is essential for the maintenance of function. As a result, comparison of DNA binding proteins of unknown function in one species with characterized DNA binding proteins in another can identify potential targets and functions. The Zn(II)2Cys6 (or C6 zinc) binuclear cluster DNA binding domain has thus far been identified exclusively in fungal proteins, generally transcriptional regulators, and there are more than 80 known or predicted proteins which contain this motif, the best characterized of which are GAL4, PPR1, LEU3, HAP1, LAC9, and PUT3. Here we review all known proteins containing the Zn(II)2Cys6 motif, along with their function, DNA binding, dimerization, and zinc(II) coordination properties and DNA binding sites. In addition, we have identified all of the Zn(II)2Cys6 motif-containing proteins in the sequence databases, including a large number with unknown function from the completed Saccharomyces cerevisiae and ongoing Schizosaccharomyces pombe genome projects, and examined the phylogenetic relationships of all the Zn(II)2Cys6 motifs from these proteins. Based on these relationships, we have assigned potential functions to a number of these unknown proteins.
- Reeves R, Nissen MS
- The A.T-DNA-binding domain of mammalian high mobility group I chromosomal proteins. A novel peptide motif for recognizing DNA structure.
- J Biol Chem. 1990; 265: 8573-82
- Display abstract
We have determined the domains of the mammalian high mobility group (HMG)I chromosomal proteins necessary and sufficient for binding to the narrow minor groove of stretches of A.T-rich DNA. Three highly conserved regions within each of the known HMG-I proteins is closely related to the consensus sequence T-P-K-R-P-R-G-R-P-K-K. A synthetic oligopeptide corresponding to this consensus "binding domain" (BD) sequence specifically binds to substrate DNA in a manner similar to the intact HMG-I proteins. Molecular Corey-Pauling-Koltun model building and computer simulations employing energy minimization programs to predict structure suggest that the consensus BD peptide has a secondary structure similar to the antitumor and antiviral drugs netropsin and distamycin and to the dye Hoechst 33258. In vitro these ligands, which also preferentially bind to A.T-rich DNA, have been demonstrated to effectively compete with both the BD peptide and the HMG-I proteins for DNA binding. The BD peptide also contains novel structural features such as a predicted Asx bend or "hook" at its amino-terminal end and laterally projecting cationic Arg/Lys side chains or "bristles" which may contribute to the binding properties of the HMG-I proteins. The predicted BD peptide structure, which we refer to as the "A.T-hook," represents a previously undescribed DNA-binding motif capable of binding to the minor groove of stretches of A.T base pairs.
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
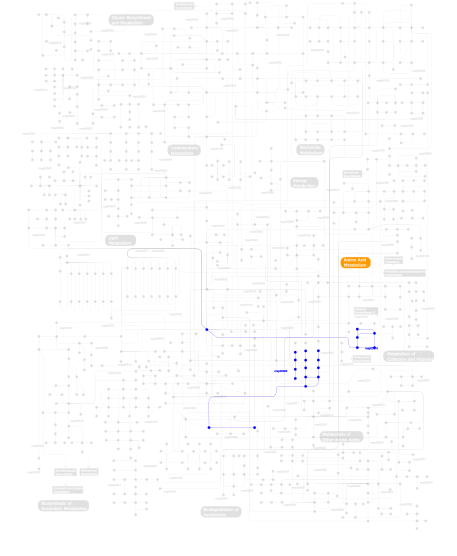
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 20.00 map04111 Cell cycle - yeast 17.14 map04530 Tight junction 17.14 map04110 Cell cycle 5.71  map00280
map00280Valine, leucine and isoleucine degradation 5.71 map04742 Taste transduction 5.71 map04010 MAPK signaling pathway 5.71 map04020 Calcium signaling pathway 5.71 map04930 Type II diabetes mellitus 5.71 map04730 Long-term depression 2.86 map03020 RNA polymerase 2.86 map03022 Basal transcription factors 2.86 map03030 DNA replication 2.86  map00310
map00310Lysine degradation This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with AT_hook domain which could be assigned to a KEGG orthologous group, and not all proteins containing AT_hook domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of AT_hook domains in PDB
PDB code Main view Title 2ezd 
SOLUTION STRUCTURE OF A COMPLEX OF THE SECOND DNA BINDING DOMAIN OF HUMAN HMG-I(Y) BOUND TO DNA DODECAMER CONTAINING THE PRDII SITE OF THE INTERFERON-BETA PROMOTER, NMR, MINIMIZED AVERAGE STRUCTURE 2eze 
SOLUTION STRUCTURE OF A COMPLEX OF THE SECOND DNA BINDING DOMAIN OF HUMAN HMG-I(Y) BOUND TO DNA DODECAMER CONTAINING THE PRDII SITE OF THE INTERFERON-BETA PROMOTER, NMR, 35 STRUCTURES 3sx0 
Crystal structure of Dot1l in complex with a brominated SAH analog 3uwp 
Crystal structure of Dot1l in complex with 5-iodotubercidin 4eqz 
Crystal structure of human DOT1L in complex with inhibitor FED2 4er0 
Crystal Structure of human DOT1L in complex with inhibitor FED1 4er7 
Crystal Structure of human DOT1L in complex with inhibitor SGC0947 4gur 
Crystal structure of LSD2-NPAC with H3 in space group P21 4gus 
Crystal structure of LSD2-NPAC with H3 in space group P3221 4gut 
Crystal structure of LSD2-NPAC 4guu 
Crystal structure of LSD2-NPAC with tranylcypromine 4hsu 
Crystal structure of LSD2-NPAC with H3(1-26)in space group P21 5juw 
5JUW - Links (links to other resources describing this domain)
-
INTERPRO IPR017956

