The domain within your query sequence starts at position and ends at position ; the E-value for the SAM domain shown below is < 1e-12.
dummy
SAMSterile alpha motif. |
|---|
| SMART accession number: | SM00454 |
|---|---|
| Description: | Widespread domain in signalling and nuclear proteins. In EPH-related tyrosine kinases, appears to mediate cell-cell initiated signal transduction via the binding of SH2-containing proteins to a conserved tyrosine that is phosphorylated. In many cases mediates homodimerisation. |
| Interpro abstract (IPR001660): | The sterile alpha motif (SAM) domain is a putative protein interaction module present in a wide variety of proteins [ (PUBMED:9007998) ] involved in many biological processes. The SAM domain that spreads over around 70 residues is found in diverse eukaryotic organisms [ (PUBMED:9886291) ]. SAM domains have been shown to homo- and hetero-oligomerise, forming multiple self-association architectures and also binding to various non-SAM domain-containing proteins [ (PUBMED:9343432) ], nevertheless with a low affinity constant [ (PUBMED:9933164) ]. SAM domains also appear to possess the ability to bind RNA [ (PUBMED:14659692) ]. Smaug, a protein that helps to establish a morphogen gradient in Drosophila embryos by repressing the translation of nanos (nos) mRNA, binds to the 3' untranslated region (UTR) of nos mRNA via two similar hairpin structures. The 3D crystal structure of the Smaug RNA-binding region shows a cluster of positively charged residues on the Smaug-SAM domain, which could be the RNA-binding surface. This electropositive potential is unique among all previously determined SAM-domain structures and is conserved among Smaug-SAM homologues. These results suggest that the SAM domain might have a primary role in RNA binding. Structural analyses show that the SAM domain is arranged in a small five-helix bundle with two large interfaces [ (PUBMED:9343432) ]. In the case of the SAM domain of EphB2, each of these interfaces is able to form dimers. The presence of these two distinct intermonomers binding surface suggest that SAM could form extended polymeric structures [ (PUBMED:9933164) ]. |
| GO function: | protein binding (GO:0005515) |
| Family alignment: |
There are 64647 SAM domains in 49933 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing SAM domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with SAM domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing SAM domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Cellular role: signalling
Binding / catalysis: protein-binding, homodimerization, SH2-binding - Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Aviv T, Lin Z, Lau S, Rendl LM, Sicheri F, Smibert CA
- The RNA-binding SAM domain of Smaug defines a new family of post-transcriptional regulators.
- Nat Struct Biol. 2003; 10: 614-621
- Display abstract
Anteroposterior patterning in Drosophila melanogaster is dependent on the sequence-specific RNA-binding protein Smaug, which binds to and regulates the translation of nanos (nos) mRNA. Here we demonstrate that the sterile-alpha motif (SAM) domain of Smaug functions as an RNA-recognition domain. This represents a new function for the SAM domain family, which is well characterized for mediating protein-protein interactions. Using homology modeling and site-directed mutagenesis, we have localized the RNA-binding surface of the Smaug SAM domain and have elaborated the RNA consensus sequence required for binding. Residues that compose the RNA-binding surface are conserved in a subgroup of SAM domain-containing proteins, suggesting that the function of the domain is conserved from yeast to humans. We show here that the SAM domain of Saccharomyces cerevisiae Vts1 binds RNA with the same specificity as Smaug and that Vts1 induces transcript degradation through a mechanism involving the cytoplasmic deadenylase CCR4. Together, these results suggest that Smaug and Vts1 define a larger class of post-transcriptional regulators that act in part through a common transcript-recognition mechanism.
- Green JB, Gardner CD, Wharton RP, Aggarwal AK
- RNA recognition via the SAM domain of Smaug.
- Mol Cell. 2003; 11: 1537-48
- Display abstract
The Nanos protein gradient in Drosophila, required for proper abdominal segmentation, is generated in part via translational repression of its mRNA by Smaug. We report here the crystal structure of the Smaug RNA binding domain, which shows no sequence homology to any previously characterized RNA binding motif. The structure reveals an unusual makeup in which a SAM domain, a common protein-protein interaction module, is affixed to a pseudo-HEAT repeat analogous topology (PHAT) domain. Unexpectedly, we find through a combination of structural and genetic analysis that it is primarily the SAM domain that interacts specifically with the appropriate nanos mRNA regulatory sequence. Therefore, in addition to their previously characterized roles in protein-protein interactions, some SAM domains play crucial roles in RNA binding.
- Williams RT, Senior PV, Van Stekelenburg L, Layton JE, Smith PJ, Dziadek MA
- Stromal interaction molecule 1 (STIM1), a transmembrane protein with growth suppressor activity, contains an extracellular SAM domain modified by N-linked glycosylation.
- Biochim Biophys Acta. 2002; 1596: 131-7
- Display abstract
Stromal interaction molecule 1 (STIM1) is a cell surface transmembrane glycoprotein implicated in tumour growth control and stromal-haematopoietic cell interactions. A single sterile alpha motif (SAM) protein-protein interaction domain is modelled within its extracellular region, a subcellular localisation not previously described for other SAM domain-containing proteins. We have defined the transmembrane topology of STIM1 by determining the sites of N-linked glycosylation. We have confirmed that STIM1 is modified by N-linked glycosylation at two sites within the SAM domain itself, deduced as asparagine residues N131 and N171, demonstrating that STIM1 is translocated across the membrane of the endoplasmic reticulum such that the SAM domain resides within the endoplasmic reticulum (ER) lumen. Both N-linked oligosaccharides remain endoglycosidase H-sensitive, indicating absence of full processing within the ER and Golgi. This immature modification is nevertheless sufficient and critical for cell surface expression of STIM1. We show that STIM1-STIM1 homotypic interactions are mediated via the cytoplasmic rather than the extracellular region of STIM1, excluding an essential role for the SAM domain in these protein interactions. These studies provide the first evidence for an extracellular localisation of a SAM domain within any protein, and the first example of a SAM domain modified by N-linked glycosylation.
- Newlon MG et al.
- The molecular basis for protein kinase A anchoring revealed by solution NMR.
- Nat Struct Biol. 1999; 6: 222-7
- Display abstract
Compartmentalization of signal transduction enzymes into signaling complexes is an important mechanism to ensure the specificity of intracellular events. Formation of these complexes is mediated by specialized protein motifs that participate in protein-protein interactions. The adenosine 3',5'-cyclic monophosphate (cAMP)-dependent protein kinase (PKA) is localized through interaction of the regulatory (R) subunit dimer with A-kinase-anchoring proteins (AKAPs). We now report the solution structure of the type II PKA R-subunit fragment RIIalpha(1-44), which encompasses both the AKAP-binding and dimerization interfaces. This structure incorporates an X-type four-helix bundle dimerization motif with an extended hydrophobic face that is necessary for high-affinity AKAP binding. NMR data on the complex between RIIalpha(1-44) and an AKAP fragment reveals extensive contacts between the two proteins. Interestingly, this same dimerization motif is present in other signaling molecules, the S100 family. Therefore, the X-type four-helix bundle may represent a conserved fold for protein-protein interactions in signal transduction.
- Smalla M et al.
- Solution structure of the receptor tyrosine kinase EphB2 SAM domain and identification of two distinct homotypic interaction sites.
- Protein Sci. 1999; 8: 1954-61
- Display abstract
The sterile alpha motif (SAM) is a protein interaction domain of around 70 amino acids present predominantly in the N- and C-termini of more than 60 diverse proteins that participate in signal transduction and transcriptional repression. SAM domains have been shown to homo- and hetero-oligomerize and to mediate specific protein-protein interactions. A highly conserved subclass of SAM domains is present at the intracellular C-terminus of more than 40 Eph receptor tyrosine kinases that are involved in the control of axonal pathfinding upon ephrin-induced oligomerization and activation in the event of cell-cell contacts. These SAM domains appear to participate in downstream signaling events via interactions with cytosolic proteins. We determined the solution structure of the EphB2 receptor SAM domain and studied its association behavior. The structure consists of five helices forming a compact structure without binding pockets or exposed conserved aromatic residues. Concentration-dependent chemical shift changes of NMR signals reveal two distinct well-separated areas on the domains' surface sensitive to the formation of homotypic oligomers in solution. These findings are supported by analytical ultracentrifugation studies. The conserved Tyr932, which was reported to be essential for the interaction with SH2 domains after phosphorylation, is buried in the hydrophobic core of the structure. The weak capability of the isolated EphB2 receptor SAM domain to form oligomers is supposed to be relevant in vivo when the driving force of ligand binding induces receptor oligomerization. A formation of SAM tetramers is thought to provide an appropriate contact area for the binding of a low-molecular-weight phosphotyrosine phosphatase and to initiate further downstream responses.
- Stapleton D, Balan I, Pawson T, Sicheri F
- The crystal structure of an Eph receptor SAM domain reveals a mechanism for modular dimerization.
- Nat Struct Biol. 1999; 6: 44-9
- Display abstract
The sterile alpha motif (SAM) domain is a novel protein module of approximately 70 amino acids that is found in a variety of signaling molecules including tyrosine and serine/threonine protein kinases, cytoplasmic scaffolding and adaptor proteins, regulators of lipid metabolism, and GTPases as well as members of the ETS family of transcription factors. The SAM domain can potentially function as a protein interaction module through the ability to homo- and hetero-oligomerize with other SAM domains. This functional property elicits the oncogenic activation of chimeric proteins arising from translocation of the SAM domain of TEL to coding regions of the betaPDGF receptor, Abl, JAK2 protein kinase and the AML1 transcription factor. Here we describe the 2.0 A X-ray crystal structure of a SAM domain homodimer from the intracellular region of the EphA4 receptor tyrosine kinase. The structure reveals a mode of dimerization that we predict is shared amongst the SAM domains of the Eph receptor tyrosine kinases and possibly other SAM domain containing proteins. These data indicate a mechanism through which an independently folding protein module can form homophilic complexes that regulate signaling events at the membrane and in the nucleus.
- Thanos CD, Goodwill KE, Bowie JU
- Oligomeric structure of the human EphB2 receptor SAM domain.
- Science. 1999; 283: 833-6
- Display abstract
The sterile alpha motif (SAM) domain is a protein interaction module that is present in diverse signal-transducing proteins. SAM domains are known to form homo- and hetero-oligomers. The crystal structure of the SAM domain from an Eph receptor tyrosine kinase, EphB2, reveals two large interfaces. In one interface, adjacent monomers exchange amino-terminal peptides that insert into a hydrophobic groove on each neighbor. A second interface is composed of the carboxyl-terminal helix and a nearby loop. A possible oligomer, constructed from a combination of these binding modes, may provide a platform for the formation of larger protein complexes.
- Kyba M, Brock HW
- The SAM domain of polyhomeotic, RAE28, and scm mediates specific interactions through conserved residues.
- Dev Genet. 1998; 22: 74-84
- Display abstract
The SAM (sterile alpha motif) domain is a 65- to 70-amino acid sequence found in many diverse proteins whose functions range from signal transduction to transcriptional repression. We show that the SAM domain of the Drosophila Polycomb group protein, polyhomeotic (ph), is capable of binding to itself in vitro. We test a number of near relatives of the ph SAM domain from fruit fly, mouse, and yeast and show that all are capable of self-binding. Heterologous interactions are seen among a subset of SAM domains, including ph, Scm, and RAE28. Several conserved amino acid residues were mutated in the ph SAM domain, and the effects on self-binding and heterologous association were demonstrated. L33, L41, and 162 are shown to be important determinants of the binding interface, while W1 and G50 are likely essential for the structure of the domain.
- Wharton RP, Sonoda J, Lee T, Patterson M, Murata Y
- The Pumilio RNA-binding domain is also a translational regulator.
- Mol Cell. 1998; 1: 863-72
- Display abstract
Posterior patterning in the Drosophila embryo requires the action of Nanos (Nos) and Pumilio (Pum), which collaborate to regulate the translation of maternal hunchback (hb) mRNA. Previous work demonstrated that Pum recognizes sites in the 3' UTR of hb mRNA. In this report, we first define the RNA-binding domain of Pum and then show that residues essential for translational repression are embedded within this domain. We also show that Nos and Pum can repress cap-independent translation from an internal ribosome entry site (IRES) in vivo, suggesting that they act downstream of the initial steps of normal, cap-dependent translation.
- Curtis D, Treiber DK, Tao F, Zamore PD, Williamson JR, Lehmann R
- A CCHC metal-binding domain in Nanos is essential for translational regulation.
- EMBO J. 1997; 16: 834-43
- Display abstract
The Drosophila Nanos protein is a localized repressor of hunchback mRNA translation in the early embryo, and is required for the establishment of the anterior-posterior body axis. Analysis of nanos mutants reveals that a small, evolutionarily conserved, C-terminal region is essential for Nanos function in vivo, while no other single portion of the Nanos protein is absolutely required. Within the C-terminal region are two unusual Cys-Cys-His-Cys (CCHC) motifs that are potential zinc-binding sites. Using absorption spectroscopy and NMR we demonstrate that the CCHC motifs each bind one equivalent of zinc with high affinity. nanos mutations disrupting metal binding at either of these two sites in vitro abolish Nanos translational repression activity in vivo. We show that full-length and C-terminal Nanos proteins bind to RNA in vitro with high affinity, but with little sequence specificity. Mutations affecting the hunchback mRNA target sites for Nanos-dependent translational repression were found to disrupt translational repression in vivo, but had little effect on Nanos RNA binding in vitro. Thus, the Nanos zinc domain does not specifically recognize target hunchback RNA sequences, but might interact with RNA in the context of a larger ribonucleoprotein complex.
- Schultz J, Ponting CP, Hofmann K, Bork P
- SAM as a protein interaction domain involved in developmental regulation.
- Protein Sci. 1997; 6: 249-53
- Display abstract
More than 60 previously undetected SAM domain-containing proteins have been identified using profile searching methods. Among these are over 40 EPH-related receptor tyrosine kinases (RPTK), Drosophila bicaudal-C, a p53 from Loligo forbesi, and diacyglycerol-kinase isoform delta. This extended dataset suggests that SAM is an evolutionary conserved protein binding domain that is involved in the regulation of numerous developmental processes among diverse eukaryotes. A conserved tyrosine in the SAM sequences of the EPH related RPTKs is likely to mediate cell-cell initiated signal transduction via the binding of SH2 containing proteins to phosphotyrosine.
- Zamore PD, Williamson JR, Lehmann R
- The Pumilio protein binds RNA through a conserved domain that defines a new class of RNA-binding proteins.
- RNA. 1997; 3: 1421-33
- Display abstract
Translation of hunchback(mat) (hb[mat]) mRNA must be repressed in the posterior of the pre-blastoderm Drosophila embryo to permit formation of abdominal segments. This translational repression requires two copies of the Nanos Response Element (NRE), a 16-nt sequence in the hb[mat] 3' untranslated region. Translational repression also requires the action of two proteins: Pumilio (PUM), a sequence-specific RNA-binding protein; and Nanos, a protein that determines the location of repression. Binding of PUM to the NRE is thought to target hb(mat) mRNA for repression. Here, we show the RNA-binding domain of PUM to be an evolutionarily conserved, 334-amino acid region at the carboxy-terminus of the approximately 158-kDa PUM protein. This contiguous region of PUM retains the RNA-binding specificity of full-length PUM protein. Proteins with sequences homologous to the PUM RNA-binding domain are found in animals, plants, and fungi. The high degree of sequence conservation of the PUM RNA-binding domain in other far-flung species suggests that the domain is an ancient protein motif, and we show that conservation of sequence reflects conservation of function: that is, the homologous region from a human protein binds RNA with sequence specificity related to but distinct from Drosophila PUM.
- Mohammadi M, Schlessinger J, Hubbard SR
- Structure of the FGF receptor tyrosine kinase domain reveals a novel autoinhibitory mechanism.
- Cell. 1996; 86: 577-87
- Display abstract
The crystal structure of the tyrosine kinase domain of fibroblast growth factor receptor 1 (FGFR1K) has been determined in its unliganded form to 2.0 angstroms resolution and in complex with with an ATP analog to 2.3 angstrosms A resolution. Several features distinguish the structure of FGFR1K from that of the tyrosine kinase domain of the insulin receptor. Residues in the activation loop of FGFR1K appear to interfere with substrate peptide binding but not with ATP binding, revealing a second and perhaps more general autoinhibitory mechanism for receptor tyrosine kinases. In addition, a dimeric form of FGFR1K observed in the crystal structure may provide insights into the molecular mechanisms by which FGF receptors are activated. Finally, the structure provides a basis for rationalizing the effects of kinase mutations in FGF receptors that lead to developmental disorders in nematodes and humans.
- Ponting CP
- SAM: a novel motif in yeast sterile and Drosophila polyhomeotic proteins.
- Protein Sci. 1995; 4: 1928-30
- Display abstract
Single copies of an approximately 65-70 residue domain are shown to be present in the sequences of 14 eukaryotic proteins, including yeast byr2, STE11, ste4, and STE50, which are essential participants in sexual differentiation. This domain, named SAM (sterile alpha motif), appears to participate in other developmental processes because it is also present in Drosophila polyhomeotic gene product and related homologues, which are thought to regulate determination of segmental specification in early embryogenesis. Its appearance in byr2 and STE11, which are MEK kinases, and in proteins containing pleckstrain homology, src homology 3, and discs-large homologous region domains, suggests possible participation in signal transduction pathways.
- Bork P, Sudol M
- The WW domain: a signalling site in dystrophin?
- Trends Biochem Sci. 1994; 19: 531-3
- Mayer BJ, Ren R, Clark KL, Baltimore D
- A putative modular domain present in diverse signaling proteins.
- Cell. 1993; 73: 629-30
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
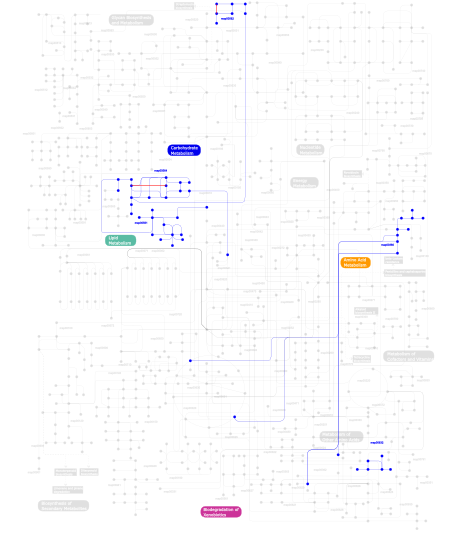
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 43.62 map04360 Axon guidance 7.98 map04070 Phosphatidylinositol signaling system 4.79  map00564
map00564Glycerophospholipid metabolism 4.79  map00561
map00561Glycerolipid metabolism 4.26 map04664 Fc epsilon RI signaling pathway 4.26  map00562
map00562Inositol phosphate metabolism 4.26 map04010 MAPK signaling pathway 4.26 map04660 T cell receptor signaling pathway 4.26 map04650 Natural killer cell mediated cytotoxicity 3.72 map04530 Tight junction 3.72 map04115 p53 signaling pathway 2.66  map00380
map00380Tryptophan metabolism 1.06  map00632
map00632Benzoate degradation via CoA ligation 0.53 map04912 GnRH signaling pathway 0.53 map04210 Apoptosis 0.53 map04740 Olfactory transduction 0.53 map04910 Insulin signaling pathway 0.53 map04742 Taste transduction 0.53 map04310 Wnt signaling pathway 0.53 map04540 Gap junction 0.53 map04020 Calcium signaling pathway 0.53 map04720 Long-term potentiation 0.53 map04916 Melanogenesis 0.53 map04340 Hedgehog signaling pathway 0.53 map04914 Progesterone-mediated oocyte maturation This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with SAM domain which could be assigned to a KEGG orthologous group, and not all proteins containing SAM domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of SAM domains in PDB
PDB code Main view Title 1b0x 
THE CRYSTAL STRUCTURE OF AN EPH RECEPTOR SAM DOMAIN REVEALS A MECHANISM FOR MODULAR DIMERIZATION. 1b4f 
OLIGOMERIC STRUCTURE OF THE HUMAN EPHB2 RECEPTOR SAM DOMAIN 1cok 
STRUCTURE OF THE C-TERMINAL DOMAIN OF P73 1dxs 
Crystal structure of the C-terminal sterile alpha motif (SAM) domain of human p73 alpha splice variant 1f0m 
MONOMERIC STRUCTURE OF THE HUMAN EPHB2 SAM (STERILE ALPHA MOTIF) DOMAIN 1kw4 
Polyhomeotic SAM domain structure 1ow5 
NMR structure of the Saccharomyces cerevisiae SAM (Sterile Alpha Motif) domain 1oxj 
Crystal structure of the Smaug RNA binding domain 1pk1 
Hetero SAM domain structure of Ph and Scm. 1pk3 
Scm SAM domain 1rg6 
Solution structure of the C-terminal domain of p63 1sgg 
THE SOLUTION STRUCTURE OF SAM DOMAIN FROM THE RECEPTOR TYROSINE KINASE EPHB2, NMR, 10 STRUCTURES 1ucv 
Sterile alpha motif (SAM) domain of ephrin type-A receptor 8 1v38 
Solution structure of the Sterile Alpha Motif (SAM) domain of mouse SAMSN1 1v85 
Sterile alpha motif (SAM) domain of mouse bifunctional apoptosis regulator 1wwv 
Solution structure of the SAM domain of human connector enhancer of KSR-like protein CNK1 1x40 
Solution structure of the SAM domain of human ARAP2 1x9x 
Solution Structure of Dimeric SAM Domain from MAPKKK Ste11 2b6g 
RNA recognition by the Vts1 SAM domain 2d3d 
crystal structure of the RNA binding SAM domain of saccharomyces cerevisiae Vts1 2d8c 
Solution structure of the sam-domain of mouse phosphatidyl ceramidecholinephosphotransferase 1 2dky 
Solution structure of the SAM-domain of Rho-GTPase-activating protein 7 2dkz 
Solution structure of the SAM_PNT-domain of the hypothetical protein LOC64762 2dl0 
Solution structure of the SAM-domain of the SAM and SH3 domain containing protein 1 2e8n 
Solution structure of the C-terminal SAM-domain of EphaA2: Ephrin type-A receptor 2 precursor (EC 2.7.10.1) 2e8o 
Solution structure of the N-terminal SAM-domain of the SAM domain and HD domain containing protein 1 (Dendritic cell-derived IFNG-induced protein) (DCIP) (Monocyte protein 5) (MOP-5) 2eam 
Solution structure of the N-terminal SAM-domain of a human putative 47 kDa protein 2ean 
Solution structure of the N-terminal SAM-domain of human KIAA0902 protein (connector enhancer of kinase suppressor of ras 2) 2eao 
Solution structure of the C-terminal SAM-domain of mouse ephrin type-B receptor 1 precursor (EC 2.7.1.112) 2eap 
Solution structure of the N-terminal SAM-domain of human lymphocyte cytosolic protein 2 2es6 
Structure of the SAM domain of Vts1p 2ese 
Structure of the SAM domain of Vts1p in complex with RNA 2f3n 
Crystal Structure of the native Shank SAM domain. 2f44 
Crystal Structure of the Zinc-bound Shank SAM domain 2f8k 
Sequence specific recognition of RNA hairpins by the SAM domain of Vts1 2fe9 
Solution structure of the Vts1 SAM domain in the presence of RNA 2gle 
Solution structure of neurabin SAM domain 2k4p 
Solution Structure of Ship2-Sam 2k60 
NMR structure of calcium-loaded STIM1 EF-SAM 2ke7 
NMR structure of the first SAM domain from AIDA1 2kg5 
NMR Solution structure of ARAP3-SAM 2kiv 
AIDA-1 SAM domain tandem 2kso 
EphA2:SHIP2 SAM:SAM complex 2l5y 
NMR structure of calcium-loaded STIM2 EF-SAM. 2lmr 
Solution structure of the first sam domain of odin 2qkq 
Structure of the SAM Domain of Human Ephrin Type-B Receptor 4 2y9t 
Structural basis of p63a SAM domain mutants involved in AEC syndrome 2y9u 
Structural basis of p63a SAM domain mutants involved in AEC syndrome 3bq7 
SAM domain of Diacylglycerol Kinase delta1 (E35G) 3bs5 
Crystal Structure of hCNK2-SAM/dHYP-SAM Complex 3bs7 
Crystal structure of the Sterile Alpha Motif (SAM) domain of Hyphen/Aveugle 3h8m 
SAM domain of human ephrin type-a receptor 7 (EPHA7) 3hil 
SAM Domain of Human Ephrin Type-A Receptor 1 (EphA1) 3k1r 
Structure of harmonin NPDZ1 in complex with the SAM-PBM of Sans 3kka 
Co-crystal structure of the sam domains of EPHA1 AND EPHA2 3sei 
Crystal Structure of Caskin1 Tandem SAMs 3sen 
Structure of Caskin1 Tandem SAMs 3tac 
Crystal Structure of the Liprin-alpha/CASK complex 3tad 
Crystal Structure of the Liprin-alpha/Liprin-beta complex 4is7 
Crystal Structure of the CASKIN2 SAM Domain Tandem 4nj8 
4NJ8 4nl9 
4NL9 4pzn 
4PZN 4pzo 
4PZO 4rqm 
4RQM 4rqn 
4RQN 4z8l 
4Z8L 5aja 
5AJA 5ao0 
5AO0 5f3x 
5F3X 5j8y 
5J8Y 5jrt 
5JRT 5jti 
5JTI 5ju5 
5JU5 5kni 
5KNI 5l1m 
5L1M - Links (links to other resources describing this domain)
-
INTERPRO IPR001660 PFAM SAM

