IQShort calmodulin-binding motif containing conserved Ile and Gln residues. |
|---|
| SMART accession number: | SM00015 |
|---|---|
| Description: | Calmodulin-binding motif. |
| Interpro abstract (IPR000048): | The IQ motif is an extremely basic unit of about 23 amino acids, whose conserved core usually fits the consensus A-x(3)-I-Q-x(2)-F-R-x(4)-K-K. The IQ motif, which can be present in one or more copies, serves as a binding site for different EF-hand proteins including the essential and regulatory myosin light chains, calmodulin (CaM), and CaM-like proteins [ (PUBMED:1558751) (PUBMED:9141499) ].Many IQ motifs are protein kinase C (PKC) phosphorylation sites [ (PUBMED:1824695) (PUBMED:8424932) ]. Resolution of the 3D structure of scallop myosin has shown that the IQ motif forms a basic amphipathic helix [ (PUBMED:8127365) ]. Some proteins known to contain an IQ motif are listed below:
This entry covers the entire IQ motif. |
| GO function: | protein binding (GO:0005515) |
| Family alignment: |
There are 165951 IQ domains in 60575 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing IQ domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with IQ domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing IQ domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Binding / catalysis: Ca2+-binding
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Deloulme JC, Prichard L, Delattre O, Storm DR
- The prooncoprotein EWS binds calmodulin and is phosphorylated by protein kinase C through an IQ domain.
- J Biol Chem. 1997; 272: 27369-77
- Display abstract
A growing family of proteins is regulated by protein kinase C and calmodulin through IQ domains, a regulatory motif originally identified in neuromodulin (Alexander, K. A., Wakim, B. T., Doyle, G. S., Walsh, K. A., and Storm, D. R. (1988) J. Biol. Chem. 263, 7544-7549). Here we report that EWS, a nuclear RNA-binding prooncoprotein, contains an IQ domain, is phosphorylated by protein kinase C, and interacts with calmodulin. Interestingly, PKC phosphorylation of EWS inhibits its binding to RNA homopolymers, and conversely, RNA binding to EWS interferes with PKC phosphorylation. Several other RNA-binding proteins, including TLS/FUS and PSF, co-purify with EWS. PKC phosphorylation of these proteins also inhibits their binding to RNA in vitro. These data suggest that PKC may regulate interactions of EWS and other RNA-binding proteins with their RNA targets and that IQ domains may provide a regulatory link between Ca2+ signal transduction pathways and RNA processing.
- Rhoads AR, Friedberg F
- Sequence motifs for calmodulin recognition.
- FASEB J. 1997; 11: 331-40
- Display abstract
Calmodulin (CaM) is recognized as a major calcium sensor and orchestrator of regulatory events through its interaction with a diverse group of cellular proteins. Many investigations have focused on defining the region of interaction between CaM and its cellular targets and the action of CaM on target protein function. Because CaM can bind with high affinity to a relatively small alpha-helical region of many proteins, success in clearly defining the essential elements of CaM binding motifs seems feasible and should provide a means of identifying CaM binding proteins. Three recognition motifs for CaM interaction are discussed in the context of experimental investigations of a variety of CaM target proteins. A modified version of the IQ motif as a consensus for Ca2+-independent binding and two related motifs for Ca2+-dependent binding, termed 18-14 and 1-5-10 based on the position of conserved hydrophobic residues, are proposed. Although considerable sequence diversity is observed among the different binding regions, these three classes of recognition motifs exist for many of the known CaM binding proteins.
- Houdusse A, Silver M, Cohen C
- A model of Ca(2+)-free calmodulin binding to unconventional myosins reveals how calmodulin acts as a regulatory switch.
- Structure. 1996; 4: 1475-90
- Display abstract
BACKGROUND: In contrast to conventional muscle myosins, where two different light chains (LCs) stabilize the elongated regulatory domain (RD) region of the head portion of the molecule, unconventional myosins are a diverse group of motors in which from one to six calmodulin (CaM) subunits are bound tandemly to the RD. In both cases, the heavy chains of the RDs have special sequences called "IQ motifs' to which the LCs or CaM bind. A previously puzzling aspect of certain unconventional myosins is their unusual mode of regulation, where activation of motility occurs at low levels of Ca2+. Although the atomic structure of the conventional muscle myosin RD has been determined, no crystallographic structure of the RD of an unconventional myosin is yet available. RESULTS: We have constructed a model of vertebrate CaM bound to the first IQ motif present in the neck region of an unconventional myosin (chicken brush border myosin I), using strict binding rules derived from the crystal structure of the scallop RD. The model accounts for aspects of the regulation of many unconventional myosins where CaM is bound at low levels of Ca2+ and released or changed in conformation at high levels of Ca2+. The conformational changes as a function of Ca2+ depend not only on the precise sequence of the IQ motifs but also on the interactions between CaM molecules bound to adjacent sites on the myosin heavy chain. CONCLUSIONS: According to our model, the full versatility of CaM binding to target peptides is displayed in the regulation of unconventional myosins. At low concentrations of Ca2+, CaM binds in a manner similar to the LCs of conventional myosins. At higher Ca2+ concentrations, CaM changes conformation and acts as a switch to regulate the activity of the unconventional myosin molecules.
- Xie X et al.
- Structure of the regulatory domain of scallop myosin at 2.8 A resolution.
- Nature. 1994; 368: 306-12
- Display abstract
The regulatory domain of scallop myosin is a three-chain protein complex that switches on this motor in response to Ca2+ binding. This domain has been crystallized and the structure solved to 2.8 A resolution. Side-chain interactions link the two light chains in tandem to adjacent segments of the heavy chain bearing the IQ-sequence motif. The Ca(2+)-binding site is a novel EF-hand motif on the essential light chain and is stabilized by linkages involving the heavy chain and both light chains, accounting for the requirement of all three chains for Ca2+ binding and regulation in the intact myosin molecule.
- Mercer JA, Seperack PK, Strobel MC, Copeland NG, Jenkins NA
- Novel myosin heavy chain encoded by murine dilute coat colour locus.
- Nature. 1991; 349: 709-13
- Display abstract
Hundreds of murine dilute mutations have been identified and analysed, making dilute one of the best genetically characterized of all mammalian loci. The recessive dilute (d) coat colour mutation carried by many inbred strains of mice produces a lightening of coat colour, caused by an abnormal adendritic melanocyte morphology that results in an uneven release of pigment granules into the developing hair shaft. Most dilute alleles (dilute-lethal) also produce a neurological defect, characterized by convulsions and opisthotonus, apparent at 8-10 days of age and continuing until the death of the animal at 2-3 weeks of age. The discovery that the original dilute allele (now termed dilute-viral or dV) is the result of the integration of an ecotropic murine leukaemia provirus has allowed the cloning of genomic DNA and in this study complementary DNA, from the dilute locus. The predicted dilute amino-acid sequence indicates that dilute encodes a novel type of myosin heavy chain, with a tail, or C-terminal, region that has elements of both type II (alpha-helical coiled-coil) and type I (non-coiled-coil) myosin heavy chains. Dilute transcripts are differentially expressed in both embryonic and adult tissues and are very abundant in neurons of the central nervous system, cephalic ganglia, and spinal ganglia. These results suggest an important role for the dilute gene product in the elaboration, maintenance, or function of cellular processes of melanocytes and neurons.
- Disease (disease genes where sequence variants are found in this domain)
-
SwissProt sequences and OMIM curated human diseases associated with missense mutations within the IQ domain.
Protein Disease Unconventional myosin-VIIa (Q13402) (SMART) OMIM:276903: Usher syndrome, type 1B ; Deafness, autosomal recessive 2, neurosensory
OMIM:600060: Deafness, autosomal dominant 11, neurosensory
OMIM:601317:UNKNOWN (SMART) OMIM:160760: Cardiomyopathy, familial hypertrophic, 1
OMIM:192600: ?Central core disease, one form - Metabolism (metabolic pathways involving proteins which contain this domain)
-
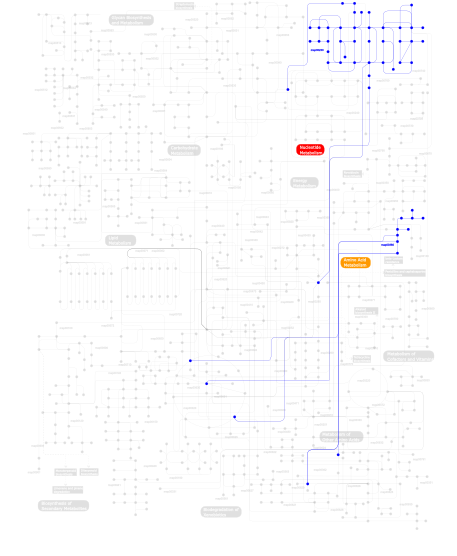
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 71.22 map04530 Tight junction 14.39 map04810 Regulation of actin cytoskeleton 5.76 map04010 MAPK signaling pathway 4.32  map00230
map00230Purine metabolism 2.88  map00380
map00380Tryptophan metabolism 1.44 map00785 Lipoic acid metabolism This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with IQ domain which could be assigned to a KEGG orthologous group, and not all proteins containing IQ domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of IQ domains in PDB
PDB code Main view Title 1b7t 
MYOSIN DIGESTED BY PAPAIN 1br1 
SMOOTH MUSCLE MYOSIN MOTOR DOMAIN-ESSENTIAL LIGHT CHAIN COMPLEX WITH MGADP.ALF4 BOUND AT THE ACTIVE SITE 1br4 
SMOOTH MUSCLE MYOSIN MOTOR DOMAIN-ESSENTIAL LIGHT CHAIN COMPLEX WITH MGADP.BEF3 BOUND AT THE ACTIVE SITE 1dfk 
NUCLEOTIDE-FREE SCALLOP MYOSIN S1-NEAR RIGOR STATE 1dfl 
SCALLOP MYOSIN S1 COMPLEXED WITH MGADP:VANADATE-TRANSITION STATE 1i84 
CRYO-EM STRUCTURE OF THE HEAVY MEROMYOSIN SUBFRAGMENT OF CHICKEN GIZZARD SMOOTH MUSCLE MYOSIN WITH REGULATORY LIGHT CHAIN IN THE DEPHOSPHORYLATED STATE. ONLY C ALPHAS PROVIDED FOR REGULATORY LIGHT CHAIN. ONLY BACKBONE ATOMS PROVIDED FOR S2 FRAGMENT. 1kk7 
SCALLOP MYOSIN IN THE NEAR RIGOR CONFORMATION 1kk8 
SCALLOP MYOSIN (S1-ADP-BeFx) IN THE ACTIN-DETACHED CONFORMATION 1kqm 
SCALLOP MYOSIN S1-AMPPNP IN THE ACTIN-DETACHED CONFORMATION 1kwo 
SCALLOP MYOSIN S1-ATPgammaS-p-PDM IN THE ACTIN-DETACHED CONFORMATION 1l2o 
SCALLOP MYOSIN S1-ADP-p-PDM IN THE ACTIN-DETACHED CONFORMATION 1m45 
CRYSTAL STRUCTURE OF MLC1P BOUND TO IQ2 OF MYO2P, A CLASS V MYOSIN 1m8q 
Molecular Models of Averaged Rigor Crossbridges from Tomograms of Insect Flight Muscle 1mvw 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1n2d 
Ternary complex of MLC1P bound to IQ2 and IQ3 of Myo2p, a class V myosin 1o18 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o19 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1a 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1b 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1c 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1d 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1e 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1f 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1o1g 
MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE 1oe9 
Crystal structure of Myosin V nucleotide-free 1qvi 
Crystal structure of scallop myosin S1 in the pre-power stroke state to 2.6 Angstrom resolution: flexibility and function in the head 1s5g 
Structure of Scallop myosin S1 reveals a novel nucleotide conformation 1scm 
STRUCTURE OF THE REGULATORY DOMAIN OF SCALLOP MYOSIN AT 2.8 ANGSTROMS RESOLUTION 1sr6 
Structure of nucleotide-free scallop myosin S1 1w7i 
Crystal Structure Of Myosin V Motor Without nucleotide soaked in 10 mM MgADP 1w7j 
Crystal Structure Of Myosin V Motor With Essential Light Chain + ADP-BeFx - Near Rigor 1wdc 
SCALLOP MYOSIN REGULATORY DOMAIN 2bki 
Myosin VI nucleotide-free (MDinsert2-IQ) crystal structure 2bl0 
Physarum polycephalum myosin II regulatory domain 2dfs 
3-D structure of Myosin-V inhibited state 2ec6 
Placopecten Striated Muscle Myosin II 2ix7 
Structure of apo-calmodulin bound to unconventional myosin V 2kxw 
Structure of the C-domain Fragment of apo Calmodulin Bound to the IQ motif of Nav1.2 2l53 
Solution NMR Structure of apo-calmodulin in complex with the IQ motif of Human Cardiac Sodium Channel NaV1.5 2m5e 
2M5E 2mys 
MYOSIN SUBFRAGMENT-1, ALPHA CARBON COORDINATES ONLY FOR THE TWO LIGHT CHAINS 2os8 
Rigor-like structures of muscle myosins reveal key mechanical elements in the transduction pathways of this allosteric motor 2otg 
Rigor-like structures of muscle myosins reveal key mechanical elements in the transduction pathways of this allosteric motor 2w4a 
Isometrically contracting insect asynchronous flight muscle 2w4g 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick stretch step 2w4h 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick release step 2w4t 
Isometrically contracting insect asynchronous flight muscle 2w4v 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick release step 2w4w 
Isometrically contracting insect asynchronous flight muscle quick frozen after a quick stretch step 3dtp 
Tarantula heavy meromyosin obtained by flexible docking to Tarantula muscle thick filament Cryo-EM 3D-MAP 3gn4 
Myosin lever arm 3i5f 
Crystal structure of squid MG.ADP myosin S1 3i5g 
Crystal structure of rigor-like squid myosin S1 3i5h 
The crystal structure of rigor like squid myosin S1 in the absence of nucleotide 3i5i 
The crystal structure of squid myosin S1 in the presence of SO4 2- 3j04 
EM structure of the heavy meromyosin subfragment of Chick smooth muscle Myosin with regulatory light chain in phosphorylated state 3jax 
3JAX 3jbh 
3JBH 3jtd 
Calcium-free Scallop Myosin Regulatory Domain with ELC-D19A Point Mutation 3jvt 
Calcium-bound Scallop Myosin Regulatory Domain (Lever Arm) with Reconstituted Complete Light Chains 3pn7 
Visualizing new hinges and a potential major source of compliance in the lever arm of myosin 3ts5 
Crystal Structure of a Light Chain Domain of Scallop Smooth Muscle Myosin 3tuy 
Phosphorylated Light Chain Domain of Scallop smooth Muscle Myosin 3wfn 
Crystal Structure of Nav1.6 IQ motif in complex with apo-CaM 4byf 
Crystal structure of human Myosin 1c in complex with calmodulin in the pre-power stroke state 4dck 
Crystal structure of the C-terminus of voltage-gated sodium channel in complex with FGF13 and CaM 4e50 
Calmodulin and Ng peptide complex 4e53 
Calmodulin and Nm peptide complex 4jpz 
Voltage-gated sodium channel 1.2 C-terminal domain in complex with FGF13U and Ca2+/calmodulin 4jq0 
Voltage-gated sodium channel 1.5 C-terminal domain in complex with FGF12B and Ca2+/calmodulin 4l79 
Crystal Structure of nucleotide-free Myosin 1b residues 1-728 with bound Calmodulin 4lzx 
Complex of IQCG and Ca2+-free CaM 4m1l 
Complex of IQCG and Ca2+-bound CaM 4ovn 
4OVN 4qbd 
4QBD 4r8g 
4R8G 4zlk 
4ZLK 5i0i 
5I0I - Links (links to other resources describing this domain)
-
INTERPRO IPR000048

