LRRLeucine-rich repeats, outliers |
|---|
| SMART accession number: | SM00370 |
|---|---|
| Description: | - |
| Family alignment: |
There are 1323917 LRR domains in 210080 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing LRR domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with LRR domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing LRR domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Cellular role: signalling, translation
Binding / catalysis: Protein-binding - Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Radauer C, Lackner P, Breiteneder H
- The Bet v 1 fold: an ancient, versatile scaffold for binding of large,hydrophobic ligands.
- BMC Evol Biol. 2008; 8: 286-286
- Display abstract
BACKGROUND: The major birch pollen allergen, Bet v 1, is a member of theubiquitous PR-10 family of plant pathogenesis-related proteins. In recentyears, a number of diverse plant proteins with low sequence similarity toBet v 1 was identified. In addition, determination of the Bet v 1structure revealed the existence of a large superfamily of structurallyrelated proteins. In this study, we aimed to identify and classify all Betv 1-related structures from the Protein Data Bank and all Bet v 1-relatedsequences from the Uniprot database. RESULTS: Structural comparisons ofrepresentative members of already known protein families structurallyrelated to Bet v 1 with all entries of the Protein Data Bank yielded 47structures with non-identical sequences. They were classified into elevenfamilies, five of which were newly identified and not included in theStructural Classification of Proteins database release 1.71. The taxonomicdistribution of these families extracted from the Pfam protein familydatabase showed that members of the polyketide cyclase family and theactivator of Hsp90 ATPase homologue 1 family were distributed among allthree superkingdoms, while members of some bacterial families wereconfined to a small number of species. Comparison of ligand bindingactivities of Bet v 1-like superfamily members revealed that theirfunctions were related to binding and metabolism of large, hydrophobiccompounds such as lipids, hormones, and antibiotics. Phylogeneticrelationships within the Bet v 1 family, defined as the group of proteinswith significant sequence similarity to Bet v 1, were determined byaligning 264 Bet v 1-related sequences. A distance-based phylogenetic treeyielded a classification into 11 subfamilies, nine exclusively containingplant sequences and two subfamilies of bacterial proteins. Plant sequencesincluded the pathogenesis-related proteins 10, the major latexproteins/ripening-related proteins subfamily, and polyketide cyclase-likesequences. CONCLUSION: The ubiquitous distribution of Bet v 1-relatedproteins among all superkingdoms suggests that a Bet v 1-like protein wasalready present in the last universal common ancestor. During evolution,this protein diversified into numerous families with low sequencesimilarity but with a common fold that succeeded as a versatile scaffoldfor binding of bulky ligands.
- Kajava AV
- Structural diversity of leucine-rich repeat proteins.
- J Mol Biol. 1998; 277: 519-27
- Display abstract
The superfamily of leucine-rich repeat proteins can be subdivided into at least six subfamilies, characterised by different lengths and consensus sequences of the repeats. It was proposed that the repeats from different subfamilies retain a similar superhelical fold, but differ in the three-dimensional structures of individual repeats. The sequence-structure relationship of three new subfamilies was examined by molecular modelling. I provide structural models for the repeats of all subfamilies. The models enable me to explain residue conservations within each subfamily. Furthermore, the difference in the packing explains why the repeats from different subfamilies never occur simultaneously in the same protein. Finally, these studies suggest different evolutionary origins for the different subfamilies. The approach used for the prediction of the leucine-rich repeat protein structures can be applied to other proteins containing internal repeats of about 20 to 30 residue in length.
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
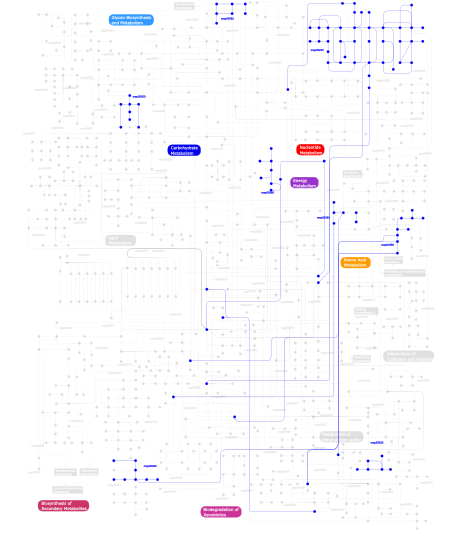
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 25.44 map04620 Toll-like receptor signaling pathway 10.25 map04080 Neuroactive ligand-receptor interaction 8.13 map04360 Axon guidance 6.71 map04120 Ubiquitin mediated proteolysis 6.36 map04512 ECM-receptor interaction 3.89 map04640 Hematopoietic cell lineage 3.89  map00230
map00230Purine metabolism 3.53 map05222 Small cell lung cancer 3.53 map04110 Cell cycle 3.18 map05120 Epithelial cell signaling in Helicobacter pylori infection 3.18 map04111 Cell cycle - yeast 2.83 map04612 Antigen processing and presentation 2.47 map04510 Focal adhesion 2.47 map04350 TGF-beta signaling pathway 2.12  map00562
map00562Inositol phosphate metabolism 2.12  map00632
map00632Benzoate degradation via CoA ligation 2.12 map04115 p53 signaling pathway 1.77  map00380
map00380Tryptophan metabolism 1.41  map00680
map00680Methane metabolism 1.41  map00940
map00940Phenylpropanoid biosynthesis 1.41  map00360
map00360Phenylalanine metabolism 0.35 map04530 Tight junction 0.35 map04010 MAPK signaling pathway 0.35 map03320 PPAR signaling pathway 0.35 map04810 Regulation of actin cytoskeleton 0.35  map00550
map00550Peptidoglycan biosynthesis This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with LRR domain which could be assigned to a KEGG orthologous group, and not all proteins containing LRR domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of LRR domains in PDB
PDB code Main view Title 1a4y 
RIBONUCLEASE INHIBITOR-ANGIOGENIN COMPLEX 1d0b 
INTERNALIN B LEUCINE RICH REPEAT DOMAIN 1dce 
CRYSTAL STRUCTURE OF RAB GERANYLGERANYLTRANSFERASE FROM RAT BRAIN 1dfj 
RIBONUCLEASE INHIBITOR COMPLEXED WITH RIBONUCLEASE A 1ds9 
SOLUTION STRUCTURE OF CHLAMYDOMONAS OUTER ARM DYNEIN LIGHT CHAIN 1 1fqv 
Insights into scf ubiquitin ligases from the structure of the skp1-skp2 complex 1fs2 
INSIGHTS INTO SCF UBIQUITIN LIGASES FROM THE STRUCTURE OF THE SKP1-SKP2 COMPLEX 1g9u 
CRYSTAL STRUCTURE OF YOPM-LEUCINE RICH EFFECTOR PROTEIN FROM YERSINIA PESTIS 1gwb 
structure of glycoprotein 1b 1h6t 
Internalin B: crystal structure of fused N-terminal domains. 1h6u 
Internalin H: crystal structure of fused N-terminal domains. 1jl5 
Novel Molecular Architecture of YopM-a Leucine-rich Effector Protein from Yersinia pestis 1k5d 
Crystal structure of Ran-GPPNHP-RanBP1-RanGAP complex 1k5g 
Crystal structure of Ran-GDP-AlFx-RanBP1-RanGAP complex 1ltx 
Structure of Rab Escort Protein-1 in complex with Rab geranylgeranyl transferase and isoprenoid 1m0z 
Crystal Structure of the von Willebrand Factor Binding Domain of Glycoprotein Ib alpha 1m10 
Crystal structure of the complex of Glycoprotein Ib alpha and the von Willebrand Factor A1 Domain 1m9l 
Relaxation-based Refined Structure Of Chlamydomonas Outer Arm Dynein Light Chain 1 1m9s 
Crystal structure of Internalin B (InlB), a Listeria monocytogenes virulence protein containing SH3-like domains. 1o6s 
Internalin (Listeria monocytogenes) / E-Cadherin (human) Recognition Complex 1o6t 
Internalin (Listeria monocytogenes) - functional domain, uncomplexed 1o6v 
Internalin (Listeria monocytogenes) - functional domain, uncomplexed 1ook 
Crystal Structure of the Complex of Platelet Receptor GPIb-alpha and Human alpha-Thrombin 1otm 
Calcium-binding mutant of the internalin B LRR domain 1otn 
Calcium-binding mutant of the Internalin B LRR domain 1oto 
Calcium-binding mutant of the internalin B LRR domain 1ozn 
1.5A Crystal Structure of the Nogo Receptor Ligand Binding Domain Reveals a Convergent Recognition Scaffold Mediating Inhibition of Myelination 1p8t 
Crystal structure of Nogo-66 Receptor 1p8v 
CRYSTAL STRUCTURE OF THE COMPLEX OF PLATELET RECEPTOR GPIB-ALPHA AND ALPHA-THROMBIN AT 2.6A 1p9a 
Crystal Structure of N-Terminal Domain of Human Platelet Receptor Glycoprotein Ib-alpha at 1.7 Angstrom Resolution 1qyy 
Crystal Structure of N-Terminal Domain of Human Platelet Receptor Glycoprotein Ib-alpha at 2.8 Angstrom Resolution 1sq0 
Crystal Structure of the Complex of the Wild-type Von Willebrand Factor A1 domain and Glycoprotein Ib alpha at 2.6 Angstrom Resolution 1u0n 
The ternary von Willebrand Factor A1-glycoprotein Ibalpha-botrocetin complex 1w8a 
Third LRR domain of Drosophila Slit 1xcd 
Dimeric bovine tissue-extracted decorin, crystal form 1 1xec 
Dimeric bovine tissue-extracted decorin, crystal form 2 1xeu 
Crystal Structure of Internalin C from Listeria monocytogenes 1xku 
Crystal structure of the dimeric protein core of decorin, the archetypal small leucine-rich repeat proteoglycan 1yrg 
THE CRYSTAL STRUCTURE OF RNA1P: A NEW FOLD FOR A GTPASE-ACTIVATING PROTEIN 1z7x 
X-ray structure of human ribonuclease inhibitor complexed with ribonuclease I 1ziw 
Human Toll-like Receptor 3 extracellular domain structure 2a0z 
The molecular structure of toll-like receptor 3 ligand binding domain 2ass 
Crystal structure of the Skp1-Skp2-Cks1 complex 2ast 
Crystal structure of Skp1-Skp2-Cks1 in complex with a p27 peptide 2bex 
Crystal structure of Placental Ribonuclease Inhibitor in complex with Human Eosinophil Derived Neurotoxin at 2A resolution. 2bnh 
PORCINE RIBONUCLEASE INHIBITOR 2ca6 
MIRAS structure determination from hemihedrally twinned crystals 2ft3 
Crystal structure of the biglycan dimer core protein 2id5 
Crystal Structure of the Lingo-1 Ectodomain 2o6q 
Structural diversity of the hagfish Variable Lymphocyte Receptors A29 2o6r 
Structural diversity of the hagfish Variable Lymphocyte Receptors B61 2o6s 
Structural diversity of the hagfish Variable Lymphocyte Receptors B59 2omt 
Crystal structure of InlA G194S+S/hEC1 complex 2omu 
Crystal structure of InlA G194S+S Y369S/hEC1 complex 2omv 
Crystal structure of InlA S192N Y369S/hEC1 complex 2omw 
Crystal structure of InlA S192N Y369S/mEC1 complex 2omx 
Crystal structure of InlA S192N G194S+S/hEC1 complex 2omy 
Crystal structure of InlA S192N/hEC1 complex 2omz 
Crystal structure of InlA Y369A/hEC1 complex 2p1m 
TIR1-ASK1 complex structure 2p1n 
Mechanism of Auxin Perception by the TIR1 Ubiqutin Ligase 2p1o 
Mechanism of Auxin Perception by the TIR1 ubiquitin ligase 2p1p 
Mechanism of Auxin Perception by the TIR1 ubiquitin ligase 2p1q 
Mechanism of Auxin Perception by the TIR1 ubiquitin ligase 2q4g 
Ensemble refinement of the protein crystal structure of human ribonuclease inhibitor complexed with ribonuclease I 2r9u 
Crystal Structure of Lamprey Variable Lymphocyte Receptor 2913 Ectodomain 2uzx 
Structure of the human receptor tyrosine kinase Met in complex with the Listeria monocytogenes invasion protein InlB: Crystal form I 2uzy 
Structure of the human receptor tyrosine kinase Met in complex with the Listeria monocytogenes invasion protein inlb: low resolution, Crystal form II 2v70 
Third LRR domain of human Slit2 2v9s 
Second LRR domain of human Slit2 2v9t 
Complex between the second LRR domain of Slit2 and The first Ig domain from Robo1 2wfh 
The Human Slit 2 Dimerization Domain D4 2wqu 
Internalin domain of Listeria monocytogenes InlB: triclinic crystal form 2wqv 
Internalin domain of Listeria monocytogenes InlB: rhombohedral crystal form 2wqw 
DOUBLE-DISULFIDE CROSS-LINKED CRYSTAL DIMER of the Listeria monocytogenes InlB internalin domain 2wqx 
InlB321_4R: S199R, D200R, G206R, A227R, C242A mutant of the Listeria monocytogenes InlB internalin domain 2xot 
Crystal structure of neuronal leucine rich repeat protein AMIGO-1 2y5q 
Listeria monocytogenes InlB (internalin B) residues 36-392 2z62 
Crystal structure of the TV3 hybrid of human TLR4 and hagfish VLRB.61 2z63 
Crystal structure of the TV8 hybrid of human TLR4 and hagfish VLRB.61 2z64 
Crystal structure of mouse TLR4 and mouse MD-2 complex 2z65 
Crystal structure of the human TLR4 TV3 hybrid-MD-2-Eritoran complex 2z66 
Crystal structure of the VT3 hybrid of human TLR4 and hagfish VLRB.61 2z7x 
Crystal structure of the TLR1-TLR2 heterodimer induced by binding of a tri-acylated lipopeptide 2z80 
Crystal structure of the TLR1-TLR2 heterodimer induced by binding of a tri-acylated lipopeptide 2z81 
Crystal structure of the TLR1-TLR2 heterodimer induced by binding of a tri-acylated lipopeptide 2z82 
Crystal structure of the TLR1-TLR2 heterodimer induced by binding of a tri-acylated lipopeptide 3a79 
Crystal structure of TLR2-TLR6-Pam2CSK4 complex 3a7b 
Crystal structure of TLR2-Streptococcus Pneumoniae lipoteichoic acid complex 3a7c 
Crystal structure of TLR2-PE-DTPA complex 3b2d 
Crystal structure of human RP105/MD-1 complex 3c6n 
Small molecule agonists and antagonists of F-box protein-substrate interactions in auxin perception and signaling 3c6o 
Small molecule agonists and antagonists of F-box protein-substrate interactions in auxin perception and signaling 3c6p 
Small molecule agonists and antagonists of F-box protein-substrate interactions in auxin perception and signaling 3cig 
Crystal structure of mouse TLR3 ectodomain 3ciy 
Mouse Toll-like receptor 3 ectodomain complexed with double-stranded RNA 3cvr 
Crystal structure of the full length IpaH3 3e6j 
Crystal Structure of Variable Lymphocyte Receptor (VLR) RBC36 in Complex with H-trisaccharide 3fxi 
Crystal structure of the human TLR4-human MD-2-E.coli LPS Ra complex 3g06 
The Salmonella Virulence Effector SspH2 Functions As A Novel E3 Ligase 3g39 
Structure of a lamprey variable lymphocyte receptor 3g3a 
Structure of a lamprey variable lymphocyte receptor in complex with a protein antigen 3g3b 
Structure of a lamprey variable lymphocyte receptor mutant in complex with a protein antigen 3j0a 
Homology model of human Toll-like receptor 5 fitted into an electron microscopy single particle reconstruction 3jbl 
3JBL 3kj4 
Structure of rat Nogo receptor bound to 1D9 antagonist antibody 3o6n 
Crystal Structure of APL1 leucine-rich repeat domain 3ogk 
Structure of COI1-ASK1 in complex with coronatine and an incomplete JAZ1 degron 3ogl 
Structure of COI1-ASK1 in complex with JA-isoleucine and the JAZ1 degron 3ogm 
Structure of COI1-ASK1 in complex with coronatine and the JAZ1 degron 3oja 
Crystal structure of LRIM1/APL1C complex 3p72 
structure of platelet Glycoprotein 1b alpha with a bound peptide inhibitor 3pmh 
Mechanism of Sulfotyrosine-Mediated Glycoprotein Ib Interaction with Two Distinct alpha-Thrombin Sites 3rfj 
Design of a binding scaffold based on variable lymphocyte receptors of jawless vertebrates by module engineering 3rfs 
Design of a binding scaffold based on variable lymphocyte receptors of jawless vertebrates by module engineering 3rg1 
Crystal structure of the RP105/MD-1 complex 3rgx 
Structural insight into brassinosteroid perception by BRI1 3rgz 
Structural insight into brassinosteroid perception by BRI1 3riz 
Crystal structure of the plant steroid receptor BRI1 ectodomain 3rj0 
Plant steroid receptor BRI1 ectodomain in complex with brassinolide 3t6q 
Crystal structure of mouse RP105/MD-1 complex 3tsr 
X-ray structure of mouse ribonuclease inhibitor complexed with mouse ribonuclease 1 3twi 
Variable Lymphocyte Receptor Recognition of the Immunodominant Glycoprotein of Bacillus anthracis Spores 3ul7 
Crystal structure of the TV3 mutant F63W 3ul8 
Crystal structure of the TV3 mutant V134L 3ul9 
structure of the TV3 mutant M41E 3ula 
Crystal structure of the TV3 mutant F63W-MD-2-Eritoran complex 3ulu 
Structure of quaternary complex of human TLR3ecd with three Fabs (Form1) 3ulv 
Structure of quaternary complex of human TLR3ecd with three Fabs (Form2) 3un9 
Crystal structure of an immune receptor 3v44 
Crystal structure of the N-terminal fragment of zebrafish TLR5 3v47 
Crystal structure of the N-tetminal fragment of zebrafish TLR5 in complex with Salmonella flagellin 3vq1 
Crystal structure of mouse TLR4/MD-2/lipid IVa complex 3vq2 
Crystal structure of mouse TLR4/MD-2/LPS complex 3w3g 
Crystal structure of human TLR8 (unliganded form) 3w3j 
Crystal structure of human TLR8 in complex with CL097 3w3k 
Crystal structure of human TLR8 in complex with CL075 3w3l 
Crystal structure of human TLR8 in complex with Resiquimod (R848) crystal form 1 3w3m 
Crystal structure of human TLR8 in complex with Resiquimod (R848) crystal form 2 3w3n 
Crystal structure of human TLR8 in complex with Resiquimod (R848) crystal form 3 3wn4 
Crystal structure of human TLR8 in complex with DS-877 3wo9 
Crystal structure of the lamprey variable lymphocyte receptor C 3wpb 
3WPB 3wpc 
3WPC 3wpd 
3WPD 3wpe 
3WPE 3wpf 
3WPF 3wpg 
3WPG 3wph 
3WPH 3wpi 
3WPI 3zyi 
NetrinG2 in complex with NGL2 3zyj 
NetrinG1 in complex with NGL1 3zyn 
Crystal structure of the N-terminal leucine rich repeats of Netrin-G Ligand-3 3zyo 
Crystal structure of the N-terminal leucine rich repeats and immunoglobulin domain of netrin-G ligand-3 4arn 
Crystal structure of the N-terminal domain of Drosophila Toll receptor 4arr 
Crystal structure of the N-terminal domain of Drosophila Toll receptor with the magic triangle I3C 4aw4 
Engineered variant of Listeria monocytogenes InlB internalin domain with an additional leucine rich repeat inserted 4b8c 
nuclease module of the yeast Ccr4-Not complex 4bsr 
Structure of the ectodomain of LGR5 in complex with R-spondin-1 ( Fu1Fu2) in P22121 crystal form 4bss 
Structure of the ectodomain of LGR5 in complex with R-spondin-1 ( Fu1Fu2) in P21 crystal form 4bst 
Structure of the ectodomain of LGR5 in complex with R-spondin-1 ( Fu1Fu2) in P6122 crystal form 4bsu 
Structure of the ectodomain of LGR5 in complex with R-spondin-1 ( Fu1Fu2) in C2 crystal form 4bv4 
Structure and allostery in Toll-Spatzle recognition 4c2a 
Crystal Structure of High-Affinity von Willebrand Factor A1 domain with R1306Q and I1309V Mutations in Complex with High Affinity GPIb alpha 4c2b 
Crystal Structure of High-Affinity von Willebrand Factor A1 domain with Disulfide Mutation in Complex with High Affinity GPIb alpha 4cc4 
Complex of InlC of Listeria monocytogenes and human Tuba C-terminal SH3 domain 4cil 
YopM-InlB: Hybrid leucine-rich repeat protein 4cnc 
Crystal structure of human 5T4 (Wnt-activated inhibitory factor 1, Trophoblast glycoprotein) 4cnm 
Crystal structure of human 5T4 (Wnt-activated inhibitory factor 1, Trophoblast glycoprotein) 4fcg 
Structure of the leucine-rich repeat domain of the type III effector XCV3220 (XopL) 4fho 
Crystal structure of an internalin C2 (inlC2) from Listeria monocytogenes str. 4b F2365 at 1.90 A resolution 4fmz 
Crystal structure of an internalin (inlF) from Listeria monocytogenes str. 4b F2365 at 1.91 A resolution 4g8a 
Crystal structure of human TLR4 polymorphic variant D299G and T399I in complex with MD-2 and LPS 4hq1 
Crystal structure of an LRR protein with two solenoids 4im6 
LRR domain from human NLRP1 4j0m 
Crystal structure of BRL1 (LRR) in complex with brassinolide 4j4l 
Modular evolution and design of the protein binding interface 4k17 
Crystal Structure of mouse CARMIL residues 1-668 4k5u 
Recognition of the BG-H Antigen by a Lamprey Variable Lymphocyte Receptor 4k79 
Recognition of the Thomsen-Friedenreich Antigen by a Lamprey Variable Lymphocyte Receptor 4kng 
Crystal structure of human LGR5-RSPO1-RNF43 4kt1 
Complex of R-spondin 1 with LGR4 extracellular domain 4kxf 
Crystal structure of NLRC4 reveals its autoinhibition mechanism 4li1 
Crystal Structures of Lgr4 and its complex with R-spondin1 4li2 
Crystal Structures of Lgr4 and its complex with R-spondin1 4lsa 
Crystal structure of BRI1 sud1 (Gly643Glu) bound to brassinolide 4lsx 
Plant steroid receptor ectodomain bound to brassinolide and SERK1 co-receptor ectodomain 4lxr 
Structure of the Toll - Spatzle complex, a molecular hub in Drosophila development and innate immunity 4lxs 
Structure of the Toll - Spatzle complex, a molecular hub in Drosophila development and innate immunity (glycosylated form) 4m7e 
Structural insight into BL-induced activation of the BRI1-BAK1 complex 4mn8 
Crystal structure of flg22 in complex with the FLS2 and BAK1 ectodomains 4mna 
Crystal structure of the free FLS2 ectodomains 4nkg 
Crystal structure of SspH1 LRR domain in complex PKN1 HR1b domain 4nkh 
Crystal structure of SspH1 LRR domain 4oqt 
LINGO-1/Li81 Fab complex 4ow2 
4OW2 4p8s 
Crystal structure of Nogo-receptor-2 4p91 
Crystal structure of the nogo-receptor-2 (27-330) 4peq 
4PEQ 4per 
4PER 4po4 
4PO4 4pq8 
Crystal Structure of Engineered Protein, Northeast Structural Genomics Consortium Target OR465 4psj 
Crystal Structure of Engineered Protein. Northeast Structural Genomics Consortium (NESG) Target OR464. 4puf 
4PUF 4q3g 
4Q3G 4q3i 
4Q3I 4q62 
Crystal Structure of Leucine-rich repeat- and Coiled coil-containing Protein from Legionella pneumophila 4qbz 
4QBZ 4qc0 
4QC0 4qdh 
4QDH 4qxe 
4QXE 4qxf 
4QXF 4r07 
4R07 4r08 
4R08 4r09 
4R09 4r0a 
4R0A 4r58 
4R58 4r5c 
4R5C 4r5d 
4R5D 4r6a 
4R6A 4r6f 
4R6F 4r6g 
4R6G 4r6j 
4R6J 4rca 
4RCA 4rcw 
4RCW 4tzh 
4TZH 4u06 
4U06 4u08 
4U08 4u09 
4U09 4u7l 
4U7L 4ufr 
4UFR 4ufs 
4UFS 4uip 
4UIP 4v2c 
4V2C 4v2d 
4V2D 4v2e 
4V2E 4wp6 
4WP6 4xa9 
4XA9 4xgo 
4XGO 4xm4 
4XM4 4xsq 
4XSQ 4y61 
4Y61 4yeb 
4YEB 4z0c 
4Z0C 4z5w 
4Z5W 4z61 
4Z61 4z62 
4Z62 4z63 
4Z63 4z64 
4Z64 5a5c 
5A5C 5aj2 
5AJ2 5awa 
5AWA 5awb 
5AWB 5awc 
5AWC 5awd 
5AWD 5az5 
5AZ5 5b0n 
5B0N 5b0t 
5B0T 5cmn 
5CMN 5cmp 
5CMP 5d3i 
5D3I 5ftt 
5FTT 5ftu 
5FTU 5gij 
5GIJ 5gmf 
5GMF 5gmg 
5GMG 5gmh 
5GMH 5gr9 
5GR9 5gs0 
5GS0 5gs2 
5GS2 5hdh 
5HDH 5hyw 
5HYW 5hzg 
5HZG 5ijb 
5IJB 5ijc 
5IJC 5ijd 
5IJD 5il7 
5IL7 5irl 
5IRL 5irm 
5IRM 5irn 
5IRN 5ixo 
5IXO 5ixq 
5IXQ 5ixt 
5IXT 5iyv 
5IYV 5iyx 
5IYX 5jh5 
5JH5

