PbH1Parallel beta-helix repeats |
|---|
| SMART accession number: | SM00710 |
|---|---|
| Description: | The tertiary structures of pectate lyases and rhamnogalacturonase A show a stack of parallel beta strands that are coiled into a large helix. Each coil of the helix represents a structural repeat that, in some homologues, can be recognised from sequence information alone. Conservation of asparagines might be connected with asparagine-ladders that contribute to the stability of the fold. Proteins containing these repeats most often are enzymes with polysaccharide substrates. |
| Interpro abstract (IPR006626): | The tertiary structures of pectate lyases and rhamnogalacturonase A show a stack of parallel beta strands that are coiled into a large helix. Each coil of the helix represents a structural repeat that, in some homologues, can be recognised from sequence information alone. Conservation of asparagines might be connected with asparagine-ladders that contribute to the stability of the fold. Proteins containing these repeats most often are enzymes with polysaccharide substrates [ (PUBMED:9724625) ]. |
| Family alignment: |
There are 585506 PbH1 domains in 92548 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing PbH1 domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with PbH1 domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing PbH1 domain in the selected taxonomic class.
- Cellular role (predicted cellular role)
-
Binding / catalysis: Polysaccharide hydrolysis
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Bedford MT, Leder P
- The FF domain: a novel motif that often accompanies WW domains.
- Trends Biochem Sci. 1999; 24: 264-5
- Jenkins J, Mayans O, Pickersgill R
- Structure and evolution of parallel beta-helix proteins.
- J Struct Biol. 1998; 122: 236-46
- Display abstract
Three bacterial pectate lyases, a pectin lyase from Aspergillus niger, thestructures of rhamnogalacturonase A from Aspergillus aculeatus, RGase A,and the P22-phage tailspike protein, TSP, display the right-handedparallel beta-helix architecture first seen in pectate lyase. The lyaseshave 7 complete coils while RGase A and TSP have 11 and 12, respectively.Each coil contains three beta-strands and three turn regions named PB1,T1, PB2, T2, PB3, and T3 in their order of occurrence. The lyases havehomologous sequences but RGase A and TSP do not show obvious sequencehomology either to the lyases or to each other. However, the structuralsimilarities between all these molecules are so extensive that divergencefrom a common ancestor is much more probable than convergence to the samefold. The region PB2-T2-PB3 is the best conserved region in the lyases andshows the clearest structural similarity. Not only is the pleating and thedirection of the hydrogen bonding in the sheets conserved, but so is theunusual alphaL-conformation turn between the two sheets. However, theoverall shape, the position of long loops, a conserved alpha-helix thatcovers the amino-terminal end of the parallel beta-helix and stacks ofresidues in alphaR-conformation at the start of PB1 all suggest a commonancestor. The functional similarity, that the enzymes all bindalpha-galactose containing polymers at an equivalent site involving PB1and its two flanking turn regions, further supports divergent evolution.We suggest that the stacking of the coils and the unusual nearperpendicular junction of PB2 and PB3 make the parallel beta-helix foldespecially likely to maintain similar main chain conformations duringdivergent evolution even after all vestige of similarity in primarystructure has vanished.
- Pickersgill R, Smith D, Worboys K, Jenkins J
- Crystal structure of polygalacturonase from Erwinia carotovora ssp.carotovora.
- J Biol Chem. 1998; 273: 24660-4
- Display abstract
The crystal structure of the 40-kDa endo-polygalacturonase from Erwiniacarotovora ssp. carotovora was solved by multiple isomorphous replacementand refined at 1.9 A to a conventional crystallographic R-factor of 0.198and Rfree of 0.239. This is the first structure of a polygalacturonase andcomprises a 10 turn right-handed parallel beta-helix domain with two loopregions forming a "tunnel like" substrate-binding cleft. Sequenceconservation indicates that the active site of polygalacturonase isbetween these two loop regions, and comparison of the structure ofpolygalacturonase with that of rhamnogalacturonase A from Aspergillusaculeatus enables two conserved aspartates, presumed to be catalyticresidues, to be identified. An adjacent histidine, in accord withbiochemical results, is also seen. A similarity in overall electrostaticproperties of the substrate-binding clefts of polygalacturonase andpectate lyase, which bind and cleave the same substrate, polygalacturonicacid, is also revealed.
- Rotin D
- WW (WWP) domains: from structure to function.
- Curr Top Microbiol Immunol. 1998; 228: 115-33
- Jurnak F, Yoder MD, Pickersgill R, Jenkins J
- Parallel beta-domains: a new fold in protein structures.
- Curr Opin Struct Biol. 1994; 4: 802-6
- Display abstract
A new type of structural domain, composed of parallel beta-strands foldedinto a coiled structure, has been observed in several protein structureswithin the past year. An analysis of the basic motif indicates that thereare two distinct types, with variations likely to be discovered in thefuture.
- Yoder MD, Lietzke SE, Jurnak F
- Unusual structural features in the parallel beta-helix in pectate lyases.
- Structure. 1993; 1: 241-51
- Display abstract
BACKGROUND: A new type of domain structure, an all parallel beta class,has recently been observed in two pectate lyases, PelC and PelE. Theatomic models have been analyzed to determine whether the new tertiaryfold exhibits unusual structural features. RESULTS: The polypeptidebackbone exhibits no new types of secondary structural elements. However,novel features occur in the amino acid side chain interactions. The sidechain atoms form linear stacks that include asparagine ladders, serinestacks, aliphatic stacks, and ringed-residue stacks. A new type ofbeta-sandwich between parallel beta-sheets is observed with propertiesthat are more characteristic of antiparallel beta-sheets. CONCLUSION: Ananalysis of the PelC and PelE structures, belonging to an all parallelbeta structural class, reveals novel amino acid side chain interactions, anew type of beta-sandwich and an atypical amino acid composition ofparallel beta-sheets. The findings are relevant to three-dimensionalstructural predictions.
- Yoder MD, Keen NT, Jurnak F
- New domain motif: the structure of pectate lyase C, a secreted plantvirulence factor.
- Science. 1993; 260: 1503-7
- Display abstract
Pectate lyases are secreted by pathogens and initiate soft-rot diseases inplants by cleaving polygalacturonate, a major component of the plant cellwall. The three-dimensional structure of pectate lyase C from Erwiniachrysanthemi has been solved and refined to a resolution of 2.2 angstroms.The enzyme folds into a unique motif of parallel beta strands coiled intoa large helix. Within the core, the amino acids form linear stacks andinclude a novel asparagine ladder. The sequence similarities that pectatelyases share with pectin lyases, pollen and style proteins, and tubulinssuggest that the parallel beta helix motif may occur in a broad spectrumof proteins.
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
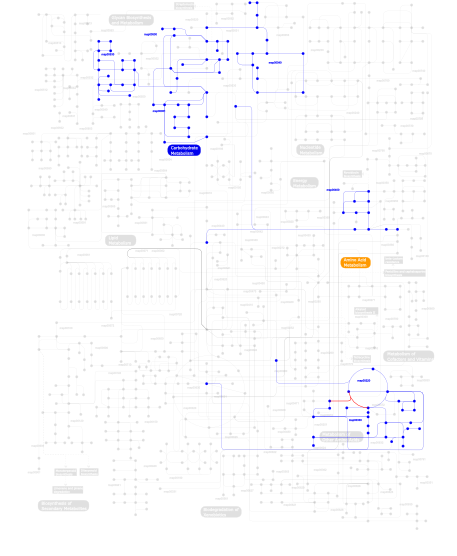
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 41.27  map00040
map00040Pentose and glucuronate interconversions 28.57  map00500
map00500Starch and sucrose metabolism 12.70  map00530
map00530Aminosugars metabolism 12.70  map00051
map00051Fructose and mannose metabolism 1.59  map00400
map00400Phenylalanine, tyrosine and tryptophan biosynthesis 1.59  map00220
map00220Urea cycle and metabolism of amino groups 1.59  map00330
map00330Arginine and proline metabolism This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with PbH1 domain which could be assigned to a KEGG orthologous group, and not all proteins containing PbH1 domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of PbH1 domains in PDB
PDB code Main view Title 1bhe 
POLYGALACTURONASE FROM ERWINIA CAROTOVORA SSP. CAROTOVORA 1clw 
TAILSPIKE PROTEIN FROM PHAGE P22, V331A MUTANT 1czf 
ENDO-POLYGALACTURONASE II FROM ASPERGILLUS NIGER 1hg8 
Endopolygalacturonase from the phytopathogenic fungus Fusarium moniliforme 1ia5 
POLYGALACTURONASE FROM ASPERGILLUS ACULEATUS 1ib4 
CRYSTAL STRUCTURE OF POLYGALACTURONASE FROM ASPERGILLUS ACULEATUS AT PH4.5 1k5c 
Endopolygalacturonase I from Stereum purpureum at 0.96 A resolution 1kcc 
Endopolygalacturonase I from Stereum purpureum complexed with a galacturonate at 1.00 A resolution. 1kcd 
Endopolygalacturonase I from Stereum purpureum complexed with two galacturonate at 1.15 A resolution. 1nhc 
Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger 1qa1 
TAILSPIKE PROTEIN, MUTANT V331G 1qa2 
TAILSPIKE PROTEIN, MUTANT A334V 1qa3 
TAILSPIKE PROTEIN, MUTANT A334I 1qq1 
TAILSPIKE PROTEIN, MUTANT E359G 1qrb 
PLASTICITY AND STERIC STRAIN IN A PARALLEL BETA-HELIX: RATIONAL MUTATIONS IN P22 TAILSPIKE PROTEIN 1rmg 
RHAMNOGALACTURONASE A FROM ASPERGILLUS ACULEATUS 1ru4 
Crystal structure of pectate lyase Pel9A 1tsp 
CRYSTAL STRUCTURE OF P22 TAILSPIKE PROTEIN: INTERDIGITATED SUBUNITS IN A THERMOSTABLE TRIMER 1tyu 
STRUCTURE OF TAILSPIKE-PROTEIN 1tyv 
STRUCTURE OF TAILSPIKE-PROTEIN 1tyw 
STRUCTURE OF TAILSPIKE-PROTEIN 1tyx 
TITLE OF TAILSPIKE-PROTEIN 1vbl 
Structure of the thermostable pectate lyase PL 47 2iq7 
Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins 2pyg 
Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module 2pyh 
Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module complexed with mannuronan trisaccharide 2v5i 
Structure of the receptor-binding protein of bacteriophage Det7: a podoviral tailspike in a myovirus 2vfm 
Low Temperature Structure of P22 Tailspike Protein Fragment (109-666) 2vfn 
Low Temperature Structure of P22 Tailspike Protein Fragment (109-666), Mutant V125A 2vfo 
Low Temperature Structure of P22 Tailspike Protein Fragment (109-666), Mutant V125L 2vfp 
Low Temperature Structure of P22 Tailspike Protein Fragment (109-666), Mutant V349L 2x3h 
COLIPHAGE K5A LYASE 2xc1 
Full-length Tailspike Protein Mutant Y108W of Bacteriophage P22 3gq7 
Crystal Structure of the Bacteriophage Phi29 Gene Product 12 N-terminal Fragment 3gq8 
Crystal Structure of the Bacteriophage phi29 gene product 12 N-terminal fragment in complex with 2-(N-cyclohexylamino)ethane sulfonic acid (CHES) 3gq9 
Crystal Structure of the Bacteriophage phi29 gene product 12 N-terminal fragment in an apo form 3gqa 
Crystal Structure of the Bacteriophage phi29 gene product 12 N-terminal fragment in complex with cobalt ions 3jur 
The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima 3suc 
Crystal structure of the pre-mature bacteriophage phi29 gene product 12 3th0 
P22 Tailspike complexed with S.Paratyphi O antigen octasaccharide 3vst 
The complex structure of XylC with Tris 3vsu 
The complex structure of XylC with xylobiose 3vsv 
The complex structure of XylC with xylose 3zpp 
Structure of the Streptococcus pneumoniae surface protein and adhesin PfbA 3zsc 
Catalytic function and substrate recognition of the pectate lyase from Thermotoga maritima 4c2l 
Crystal structure of endo-xylogalacturonan hydrolase from Aspergillus tubingensis 4hwv 
Structure of Pectate Lyase from Acidovorax avenae subsp citrulli 4mr0 
Crystal structure of PfbA, a surface adhesin of Streptococcus pneumoniae 4nk6 
Crystal Structure of the periplasmic alginate epimerase AlgG 4nk8 
Crystal Structure of the periplasmic alginate epimerase AlgG D317A mutant 4ozy 
4OZY 4ozz 
4OZZ 4rmx 
4RMX 4ru4 
4RU4 5gai 
5GAI - Links (links to other resources describing this domain)
-
INTERPRO IPR006626

