PASPAS domain |
|---|
| SMART accession number: | SM00091 |
|---|---|
| Description: | PAS motifs appear in archaea, eubacteria and eukarya. Probably the most surprising identification of a PAS domain was that in EAG-like K+-channels ([1]; Ponting & Aravind, in press). |
| Interpro abstract (IPR000014): | PAS domains are involved in many signalling proteins where they are used as a signal sensor domain [ (PUBMED:10357859) ]. PAS domains appear in archaea, bacteria and eukaryotes. Several PAS-domain proteins are known to detect their signal by way of an associated cofactor. Heme, flavin, and a 4-hydroxycinnamyl chromophore are used in different proteins. The PAS domain was named after three proteins that it occurs in:
PAS domains are often associated with PAC domains IPR001610 . It appears that these domains are directly linked, and that together they form the conserved 3D PAS fold. The division between the PAS and PAC domains is caused by major differences in sequences in the region connecting these two motifs [ (PUBMED:15009198) ]. In human PAS kinase, this region has been shown to be very flexible, and adopts different conformations depending on the bound ligand [ (PUBMED:12377121) ]. Probably the most surprising identification of a PAS domain was that in EAG-like K -channels [ (PUBMED:9301332) ]. |
| Family alignment: |
There are 613402 PAS domains in 396128 proteins in SMART's nrdb database.
Click on the following links for more information.
- Evolution (species in which this domain is found)
-
Taxonomic distribution of proteins containing PAS domain.
This tree includes only several representative species. The complete taxonomic breakdown of all proteins with PAS domain is also avaliable.
Click on the protein counts, or double click on taxonomic names to display all proteins containing PAS domain in the selected taxonomic class.
- Literature (relevant references for this domain)
-
Primary literature is listed below; Automatically-derived, secondary literature is also avaliable.
- Zhulin IB, Taylor BL, Dixon R
- PAS domain S-boxes in Archaea, Bacteria and sensors for oxygen and redox.
- Trends Biochem Sci. 1997; 22: 331-3
- Borgstahl GE, Williams DR, Getzoff ED
- 1.4 A structure of photoactive yellow protein, a cytosolic photoreceptor: unusual fold, active site, and chromophore.
- Biochemistry. 1995; 34: 6278-87
- Display abstract
A photosensing protein directs light energy captured by its chromophore into a photocycle. The protein's structure must accommodate the photocycle and promote the resulting chemical or conformational changes that lead to signal transduction. The 1.4 A crystallographic structure of photoactive yellow protein, determined by multiple isomorphous replacement methods, provides the first view at atomic resolution of a protein with a photocycle. The alpha/beta fold, which differs from the original chain tracing, shows striking similarity to distinct parts of the signal transduction proteins profilin and the SH2 domain. In the dark state structure of photoactive yellow protein, the novel 4-hydroxycinnamyl chromophore, covalently attached to Cys69, is buried within the major hydrophobic core of the protein and is tethered at both ends by hydrogen bonds. In the active site, the yellow anionic form of the chromophore is stabilized by hydrogen bonds from the side chains of Tyr42 and buried Glu46 to the phenolic oxygen atom and by electrostatic complementarity with the positively charged guanidinium group of Arg52. Thr50 further interlocks Tyr42, Glu46, and Arg52 through a network of active site hydrogen bonds. Arg52, located in a concavity of the protein surface adjacent to the dominant patch of negative electrostatic potential, shields the chromophore from solvent and is positioned to form a gateway for the phototactic signal. Overall, the high-resolution structure of photoactive yellow protein supports a mechanism whereby electrostatic interactions create an active site poised for photon-induced rearrangements and efficient protein-mediated signal transduction.
- Metabolism (metabolic pathways involving proteins which contain this domain)
-
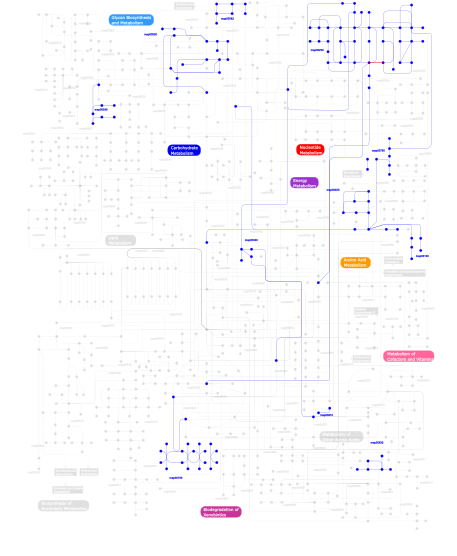
Click the image to view the interactive version of the map in iPath% proteins involved KEGG pathway ID Description 62.41 map02020 Two-component system - General 21.07 map02030 Bacterial chemotaxis - General 4.56 map04710 Circadian rhythm 3.65 map05211 Renal cell carcinoma 2.53 map03090 Type II secretion system 1.72  map00230
map00230Purine metabolism 1.32 map04914 Progesterone-mediated oocyte maturation 1.01 map04150 mTOR signaling pathway 0.20  map00562
map00562Inositol phosphate metabolism 0.20 map04540 Gap junction 0.20  map00632
map00632Benzoate degradation via CoA ligation 0.20  map00540
map00540Lipopolysaccharide biosynthesis 0.20 map04730 Long-term depression 0.10  map00910
map00910Nitrogen metabolism 0.10  map00190
map00190Oxidative phosphorylation 0.10  map00130
map00130Ubiquinone biosynthesis 0.10  map00400
map00400Phenylalanine, tyrosine and tryptophan biosynthesis 0.10  map00790
map00790Folate biosynthesis 0.10  map00340
map00340Histidine metabolism 0.10  map00500
map00500Starch and sucrose metabolism This information is based on mapping of SMART genomic protein database to KEGG orthologous groups. Percentage points are related to the number of proteins with PAS domain which could be assigned to a KEGG orthologous group, and not all proteins containing PAS domain. Please note that proteins can be included in multiple pathways, ie. the numbers above will not always add up to 100%.
- Structure (3D structures containing this domain)
3D Structures of PAS domains in PDB
PDB code Main view Title 1d06 
STRUCTURAL BASIS OF DIMERIZATION AND SENSORY MECHANISMS OF OXYGEN-SENSING DOMAIN OF RHIZOBIUM MELILOTI FIXL DETERMINED AT 1.4A RESOLUTION 1d7e 
CRYSTAL STRUCTURE OF THE P65 CRYSTAL FORM OF PHOTOACTIVE YELLOW PROTEIN 1dp6 
OXYGEN-BINDING COMPLEX OF FIXL HEME DOMAIN 1dp8 
CRYSTAL STRUCTURE OF THE NITRIC OXIDE BOUND FIXL HEME DOMAIN 1dp9 
CRYSTAL STRUCTURE OF IMIDAZOLE-BOUND FIXL HEME DOMAIN 1drm 
CRYSTAL STRUCTURE OF THE LIGAND FREE BJFIXL HEME DOMAIN 1ew0 
CRYSTAL STRUCTURE ANALYSIS OF THE SENSOR DOMAIN OF RMFIXL(FERROUS FORM) 1f98 
CRYSTAL STRUCTURE OF THE PHOTOACTIVE YELLOW PROTEIN MUTANT T50V 1f9i 
CRYSTAL STRUCTURE OF THE PHOTOACTIVE YELLOW PROTEIN MUTANT Y42F 1gsv 
CRYSTAL STRUCTURE OF THE P65 CRYSTAL FORM OF PHOTOACTIVE YELLOW PROTEIN 1gsw 
CRYSTAL STRUCTURE OF THE P65 CRYSTAL FORM OF PHOTOACTIVE YELLOW PROTEIN G51S MUTANT 1gsx 
CRYSTAL STRUCTURE OF THE P65 CRYSTAL FORM OF PHOTOACTIVE YELLOW PROTEIN G47S/G51S MUTANT 1kou 
Crystal Structure of the Photoactive Yellow Protein Reconstituted with Caffeic Acid at 1.16 A Resolution 1ll8 
Structure and interactions of PAS kinase N-terminal PAS domain: Model for intramolecular kinase regulation 1lsv 
Crystal structure of the CO-bound BjFixL heme domain 1lsw 
Crystal structure of the ferrous BjFixL heme domain 1lsx 
Crystal structure of the methylimidazole-bound BjFixL heme domain 1lt0 
Crystal structure of the CN-bound BjFixL heme domain 1mzu 
Crystal Structure of the Photoactive Yellow Protein Domain from the Sensor Histidine Kinase Ppr from Rhodospirillum centenum 1nwz 
PYP Ultra-high resolution structure of a Bacterial Photoreceptor 1odv 
Photoactive yellow protein 1-25 deletion mutant 1ot6 
CRYOTRAPPED CRYSTAL STRUCTURE OF THE E46Q MUTANT OF PHOTOACTIVE YELLOW PROTEIN UNDER CONTINUOUS ILLUMINATION AT 110K 1ot9 
CRYOTRAPPED STATE IN WILD TYPE PHOTOACTIVE YELLOW PROTEIN, INDUCED WITH CONTINUOUS ILLUMINATION AT 110K 1ota 
E46Q MUTANT OF PHOTOACTIVE YELLOW PROTEIN, P63 AT 295K 1otb 
WILD TYPE PHOTOACTIVE YELLOW PROTEIN, P63 AT 295K 1otd 
STRONG HYDROGEN BONDS IN PHOTOACTIVE YELLOW PROTEIN AND THEIR ROLE IN ITS PHOTOCYCLE 1ote 
E46Q MUTANT OF PHOTOACTIVE YELLOW PROTEIN, P65 AT 110K 1oti 
E46Q MUTANT OF PHOTOACTIVE YELLOW PROTEIN, P65 AT 295K 1p97 
NMR structure of the C-terminal PAS domain of HIF2a 1s1y 
Photoactivated chromophore conformation in Photoactive Yellow Protein (E46Q mutant) from 10 microseconds to 3 milliseconds 1s1z 
Photoactivated chromophore conformation in Photoactive Yellow Protein (E46Q mutant) from 10 to 500 nanoseconds 1s4r 
Structure of a reaction intermediate in the photocycle of PYP extracted by a SVD-driven analysis 1s4s 
Reaction Intermediate in the Photocycle of PYP, intermediate occupied between 100 micro-seconds to 5 milli-seconds 1s66 
Crystal structure of heme domain of direct oxygen sensor from E. coli 1s67 
Crystal structure of heme domain of direct oxygen sensor from E. coli 1t18 
Early intermediate IE1 from time-resolved crystallography of the E46Q mutant of PYP 1t19 
Early intermediate IE2 from time-resolved crystallography of the E46Q mutant of PYP 1t1a 
Late intermediate IL1 from time-resolved crystallography of the E46Q mutant of PYP 1t1b 
Late intermediate IL2 from time-resolved crystallography of the E46Q mutant of PYP 1t1c 
Late intermediate IL3 from time-resolved crystallography of the E46Q mutant of PYP 1ts0 
Structure of the pB1 intermediate from time-resolved Laue crystallography 1ts6 
Structure of the pB2 intermediate from time-resolved Laue crystallography 1ts7 
Structure of the pR cis wobble and pR E46Q intermediates from time-resolved Laue crystallography 1ts8 
Structure of the pR cis planar intermediate from time-resolved Laue crystallography 1ugu 
Crystal structure of PYP E46Q mutant 1uwn 
The Initial Events in the Photocycle of Photoactive Yellow Protein: A Common Mechanism on Light Activation in Photoreceptor Proteins 1uwp 
Initial Events in the Photocycle of Photoactive Yellow Protein 1v9y 
Crystal Structure of the heme PAS sensor domain of Ec DOS (ferric form) 1v9z 
Crystal Structure of the heme PAS sensor domain of Ec DOS (Ferrous Form) 1vb6 
Crystal Structure of the heme PAS sensor domain of Ec DOS (oxygen-bound form) 1wa9 
Crystal Structure of the PAS repeat region of the Drosophila clock protein PERIOD 1x0o 
human ARNT C-terminal PAS domain 1xfn 
NMR structure of the ground state of the photoactive yellow protein lacking the N-terminal part 1xfq 
structure of the blue shifted intermediate state of the photoactive yellow protein lacking the N-terminal part 1xj2 
CO-bound structure of bjFixLH 1xj3 
bjFixLH in unliganded ferrous form 1xj4 
CO-bound structure of BjFixLH 1xj6 
Structure of bjFixLH in the unliganded ferrous form 1y28 
Crystal structure of the R220A metBJFIXL HEME domain 1ztu 
Structure of the chromophore binding domain of bacterial phytochrome 2a24 
HADDOCK Structure of HIF-2a/ARNT PAS-B Heterodimer 2b02 
Crystal Structure of ARNT PAS-B Domain 2cmn 
A Proximal Arginine Residue in the Switching Mechanism of the FixL Oxygen Sensor 2d01 
Wild Type Photoactive Yellow Protein, P65 Form 2d02 
R52Q Mutant of Photoactive Yellow Protein, P65 Form 2gj3 
Crystal structure of the FAD-containing PAS domain of the protein NifL from Azotobacter vinelandii. 2hv1 
HADDOCK structure of ARNT PAS-B Homodimer 2i9v 
Structural role of Y98 in PYP: effects on fluorescence, gateway and photocycle recovery 2jhe 
N-terminal domain of TyrR transcription factor (residues 1 - 190) 2k7s 
Human ARNT C-Terminal PAS Domain, 3 Residue IB slip 2kdk 
Structure of human circadian clock protein BMAL2 C-terminal PAS domain 2kx6 
Signaling state of Photoactive Yellow Protein 2l0w 
Solution NMR structure of the N-terminal PAS domain of HERG potassium channel 2l1m 
Solution structure of the eag domain of the hERG (Kv11.1) K+ channel 2l4r 
NMR solution structure of the N-terminal PAS domain of hERG 2mwg 
2MWG 2o9b 
Crystal Structure of Bacteriophytochrome chromophore binding domain 2o9c 
Crystal Structure of Bacteriophytochrome chromophore binding domain at 1.45 angstrom resolution 2ohh 
Crystal Structure of coenzyme F420H2 oxidase (FprA), a diiron flavoprotein, active oxidized state 2ohi 
Crystal Structure of coenzyme F420H2 oxidase (FprA), a diiron flavoprotein, reduced state 2ohj 
Crystal Structure of coenzyme F420H2 oxidase (FprA), a diiron flavoprotein, inactive oxidized state 2owh 
Structure of an early-microsecond photolyzed state of CO-bjFixLH 2owj 
Structure of an early-microsecond photolyzed state of CO-bjFixLH, dark state 2phy 
PHOTOACTIVE YELLOW PROTEIN, DARK STATE (UNBLEACHED) 2pr5 
Structural Basis for Light-dependent Signaling in the Dimeric LOV Photosensor YtvA (Dark Structure) 2pr6 
Structural Basis for Light-dependent Signaling in the Dimeric LOV Photosensor YtvA (Light Structure) 2pyp 
PHOTOACTIVE YELLOW PROTEIN, PHOTOSTATIONARY STATE, 50% GROUND STATE, 50% BLEACHED 2pyr 
PHOTOACTIVE YELLOW PROTEIN, 1 NANOSECOND INTERMEDIATE (287K) 2qj5 
PYP ultra-high resolution of a bacterial photoreceptor 2qj7 
PYP ultra-high resolution of a bacterial photoreceptor 2qws 
Neutron and X-ray structural studies of short hydrogen bonds in Photoactive Yellow Protein (PYP) 2r78 
Crystal structure of a domain of the sensory box sensor histidine kinase/response regulator from Geobacter sulfurreducens 2v0u 
n- and c-terminal helices of oat lov2 (404-546) are involved in light-induced signal transduction (cryo dark structure of lov2 (404-546)) 2v0w 
N- and C-terminal helices of oat LOV2 (404-546) are involved in light- induced signal transduction (cryo-trapped light structure of LOV2 ( 404-546)) 2v1a 
N- and C-terminal helices of oat LOV2 (404-546) are involved in light-induced signal transduction (room temperature (293K) dark structure of LOV2 (404-546)) 2v1b 
N- and C-terminal helices of oat LOV2 (404-546) are involved in light-induced signal transduction (room temperature (293K) light structure of LOV2 (404-546)) 2vea 
The complete sensory module of the cyanobacterial phytochrome Cph1 in the Pr-state. 2vlg 
KinA PAS-A domain, homodimer 2vv6 
BJFIXLH IN FERRIC FORM 2vv7 
BJFIXLH IN UNLIGANDED FERROUS FORM 2vv8 
Molecular mechanism of signal transduction in bjFixL 2w0n 
Plasticity of PAS domain and potential role for signal transduction in the histidine-kinase DcuS 2wkp 
Structure of a photoactivatable Rac1 containing Lov2 Wildtype 2wkq 
Structure of a photoactivatable Rac1 containing the Lov2 C450A Mutant 2wkr 
Structure of a photoactivatable Rac1 containing the Lov2 C450M Mutant 2z6c 
Crystal structure of LOV1 domain of phototropin1 from Arabidopsis thaliana 2z6d 
Crystal structure of LOV1 domain of phototropin2 from Arabidopsis thaliana 2zoh 
X-ray Crystal Structure of Photoactive Yellow Protein, Wild type, at 295K 2zoi 
Neutron Crystal Structure of Photoactive Yellow Protein, Wild type, at 295K 3a0r 
Crystal structure of histidine kinase ThkA (TM1359) in complex with response regulator protein TrrA (TM1360) 3a0s 
PAS domain of histidine kinase ThkA (TM1359) 3a0v 
PAS domain of histidine kinase ThkA (TM1359) (SeMet, F486M/F489M) 3b33 
Crystal structure of the PAS domain of nitrogen regulation protein NR(II) from Vibrio parahaemolyticus 3bwl 
Crystal structure of PAS domain of HTR-like protein from Haloarcula marismortui 3eeh 
The crystal structure of the domain of the putative light and redox sensing histidine kinase from Haloarcula marismortui 3ef0 
The Structure of Fcp1, an essential RNA polymerase II CTD phosphatase 3ewk 
Structure of the redox sensor domain of Methylococcus capsulatus (Bath) MmoS 3f1n 
Crystal structure of a high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains, with internally bound ethylene glycol. 3f1o 
Crystal structure of the high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains, with an internally-bound artificial ligand 3f1p 
Crystal structure of a high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains 3fc7 
The crystal structure of a domain of HTR-like protein from Haloarcula marismortui ATCC 43049 3fg8 
Crystal structure of PAS domain of RHA05790 3gdi 
Mammalian Clock Protein mPER2 - Crystal Struture of a PAS Domain Fragment 3gec 
Crystal structure of a tandem PAS domain fragment of Drosophila PERIOD 3h7w 
Crystal structure of the high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains with the artificial ligand THS017 3h82 
Crystal structure of the high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains with the artificial ligand THS020 3k3c 
The N-terminal PAS domain crystal structure of Rv1364c from Mycobacterium tuberculosis at 1.62 3k3d 
The N-terminal PAS domain crystal structure of RV1364C from Mycobacterium Tuberculosis at 2.3 angstrom 3kx0 
Crystal Structure of the PAS domain of Rv1364c 3luq 
The Crystal Structure of a PAS Domain from a Sensory Box Histidine Kinase Regulator from Geobacter sulfurreducens to 2.5A 3lyx 
Crystal structure of the PAS domain of the protein CPS_1291 from Colwellia psychrerythraea. Northeast Structural Genomics Consortium target id CsR222B 3mfx 
Crystal Structure of the sensory box domain of the sensory-box/GGDEF protein SO_1695 from Shewanella oneidensis, Northeast Structural Genomics Consortium Target SoR288B 3mjq 
Crystal structure of the PAS domain of Q24QT8_DESHY protein from Desulfitobacterium hafniense. Northeast Structural Genomics Consortium Target DhR85c. 3mqo 
The Crystal Structure of the PAS domain in complex with isopropanol of a Transcriptional Regulator in the LuxR family from Burkholderia thailandensis to 1.7A 3mqq 
The Crystal Structure of the PAS domain in complex with Ethanol of a Transcriptional Regulator in the LuxR family from Burkholderia thailandensis to 1.65A 3mr0 
Crystal Structure of Sensory Box Histidine Kinase/Response Regulator from Burkholderia thailandensis E264 3mxq 
Crystal structure of sensory box sensor histidine kinase from Vibrio cholerae 3nja 
The crystal structure of the PAS domain of a GGDEF family protein from Chromobacterium violaceum ATCC 12472. 3olo 
Crystal structure of a PAS domain from two-component sensor histidine kinase 3p7n 
Crystal structure of light activated transcription factor El222 from Erythrobacter litoralis 3phy 
PHOTOACTIVE YELLOW PROTEIN, DARK STATE (UNBLEACHED), SOLUTION STRUCTURE, NMR, 26 STRUCTURES 3pyp 
PHOTOACTIVE YELLOW PROTEIN, CRYOTRAPPED EARLY LIGHT CYCLE INTERMEDIATE 3rty 
Structure of an Enclosed Dimer Formed by The Drosophila Period Protein 3s7n 
Crystal Structure of the alternate His 207 conformation of the Infrared Fluorescent D207H variant of Deinococcus Bacteriophytochrome chromophore binding domain at 2.45 angstrom resolution 3s7o 
Crystal Structure of the Infrared Fluorescent D207H variant of Deinococcus Bacteriophytochrome chromophore binding domain at 1.24 angstrom resolution 3s7p 
Crystal Structure of the Infrared Fluorescent D207H variant of Deinococcus Bacteriophytochrome chromophore binding domain at 1.72 angstrom resolution 3s7q 
Crystal Structure of a Monomeric Infrared Fluorescent Deinococcus radiodurans Bacteriophytochrome chromophore binding domain 3sw1 
Structure of a full-length bacterial LOV protein 3ue6 
The dark structure of the blue-light photoreceptor Aureochrome1 LOV 3ulf 
The light state structure of the blue-light photoreceptor Aureochrome1 LOV 3umd 
Structure of pB intermediate of Photoactive yellow protein (PYP) at pH 4. 3ume 
Structure of pB intermediate of Photoactive yellow protein (PYP) at pH 7 3ve3 
Structure of ICT Intermediate from time-resolved laue crystallography 3ve4 
Structures of ICT and PR1 intermediates from time-resolved laue crystallography 3vol 
X-ray Crystal Structure of PAS-HAMP Aer2 in the CN-bound Form 3zq5 
Structure of the Y263F mutant of the cyanobacterial phytochrome Cph1 4b9o 
The PR0 Photocycle Intermediate of Photoactive Yellow Protein 4bbt 
The PR1 Photocycle Intermediate of Photoactive Yellow Protein 4bbu 
The PR2 Photocycle Intermediate of Photoactive Yellow Protein 4bbv 
The PB0 Photocycle Intermediate of Photoactive Yellow Protein 4cqh 
4CQH 4dj2 
Unwinding the Differences of the Mammalian PERIOD Clock Proteins from Crystal Structure to Cellular Function 4dj3 
Unwinding the Differences of the Mammalian PERIOD Clock Proteins from Crystal Structure to Cellular Function 4eq1 
Crystal Structure of the ARNT PAS-B homodimer 4ew7 
The crystal structure of conjugative transfer PAS_like domain from Salmonella enterica subsp. enterica serovar Typhimurium 4f3l 
Crystal Structure of the Heterodimeric CLOCK:BMAL1 Transcriptional Activator Complex 4gcz 
Structure of a blue-light photoreceptor 4ghi 
Crystal structure of the high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains in complex with a benzoxadiazole antagonist 4gs9 
Crystal structure of the high affinity heterodimer of HIF2 alpha and ARNT C-terminal PAS domains in complex with an inactive benzoxadiazole antagonist 4gw9 
Structure of a bacteriophytochrome and light-stimulated protomer swapping with a gene repressor 4h6j 
Identification of Cys 255 in HIF-1 as a novel site for development of covalent inhibitors of HIF-1 /ARNT PasB domain protein-protein interaction. 4hh2 
Structure of PpsR without the HTH motif from Rb. sphaeroides 4hh3 
Structure of the AppA-PpsR2 core complex from Rb. sphaeroides 4hhd 
2.75 Angstrom resolution crystal structure of the A. thaliana LOV2 domain with an extended N-terminal A' helix (cryo dark structure) 4hi4 
Crystal structure of the 5-coordinate ferric heme-binding PAS domain of Aer2 from P. aeruginosa 4hia 
Crystal Structure of Rhodobacter Sphaeroides LOV protein 4hj3 
Crystal Structure of Rhodobacter Sphaeroides LOV protein 4hj4 
Crystal Structure of Rhodobacter Sphaeroides LOV protein 4hj6 
Crystal Structure of Rhodobacter Sphaeroides LOV protein 4hnb 
Crystal Structure of Rhodobacter Sphaeroides LOV protein 4hp4 
Crystal structure of PAS domain from the fruit-fly ELK potassium channel 4hp9 
Crystal structure of the N-terminal truncated PAS domain from the hERG potassium channel 4hqa 
Crystal structure of PAS domain from the human ERG (hERG) potassium channel 4hy8 
Structures of PR1 and PR2 intermediates from time-resolved laue crystallography 4i38 
Structures of IT intermediates from time-resolved laue crystallography collected at 14ID-B, APS 4i39 
Structures of ICT and PR1 intermediates from time-resolved laue crystallography collected at 14ID-B, APS 4i3a 
Structures of PR1 and PR2 intermediates from time-resolved laue crystallography collected at 14ID-B, APS 4i3i 
Structures of IT intermediate of photoactive yellow prtein E46Q muntant from time-resolved laue crystallography collected at 14ID APS 4i3j 
Structures of PR1 intermediate of photoactive yellow prtein E46Q muntant from time-resolved laue crystallography collected AT 14ID APS 4i5s 
Structure and function of sensor histidine kinase 4ijg 
Crystal structure of monomeric bacteriophytochrome 4kqd 
4KQD 4kuk 
4KUK 4kuo 
4KUO 4l9e 
Structure of PpsR Q-PAS1 from Rb. sphaeroides 4l9f 
Structure of a SeMet derivative of PpsR Q-PAS1 from Rb. sphaeroides 4l9g 
Structure of PpsR N-Q-PAS1 from Rb. sphaeroides 4llo 
Structure of the eag domain-CNBHD complex of the mouse EAG1 channel 4lpz 
4LPZ 4lrx 
Crystal Structure of the E.coli DhaR(N)-DhaK complex 4lry 
Crystal Structure of the E.coli DhaR(N)-DhaK(T79L) complex 4lrz 
Crystal Structure of the E.coli DhaR(N)-DhaL complex 4m4x 
Structure and Dimerization Properties of the Aryl Hydrocarbon Receptor (AHR) PAS-A Domain 4mn5 
4MN5 4mn6 
4MN6 4o01 
Crystal Structure of D. radiodurans Bacteriophytochrome Photosensory Core Module in its Illuminated Form 4o0p 
Crystal Structure of D. radiodurans Bacteriophytochrome Photosensory Core Module in its Dark Form 4o8g 
4O8G 4pky 
4PKY 4q0h 
4Q0H 4q0i 
4Q0I 4q0j 
4Q0J 4r38 
4R38 4r3a 
4R3A 4rpw 
4RPW 4rq9 
4RQ9 4wf0 
4WF0 4wl9 
4WL9 4wla 
4WLA 4wn5 
4WN5 4xpz 
4XPZ 4xq0 
4XQ0 4xt2 
4XT2 4y3i 
4Y3I 4y5f 
4Y5F 4z1w 
4Z1W 4zp4 
4ZP4 4zph 
4ZPH 4zpk 
4ZPK 4zpr 
4ZPR 4zqd 
4ZQD 4zrr 
4ZRR 5a8b 
5A8B 5ajg 
5AJG 5akp 
5AKP 5c5k 
5C5K 5djt 
5DJT 5dju 
5DJU 5dkk 
5DKK 5dkl 
5DKL 5efw 
5EFW 5hd3 
5HD3 5hd5 
5HD5 5hdc 
5HDC 5hdd 
5HDD 5hds 
5HDS 5iu1 
5IU1 5j3w 
5J3W 5j4e 
5J4E 5j7e 
5J7E 5k5b 
5K5B 5k7l 
5K7L 5kiz 
5KIZ 5l8m 
5L8M 5lbr 
5LBR 5sy5 
5SY5 5sy7 
5SY7 5t0t 
5T0T 5tbm 
5TBM - Links (links to other resources describing this domain)
-
INTERPRO IPR000014

